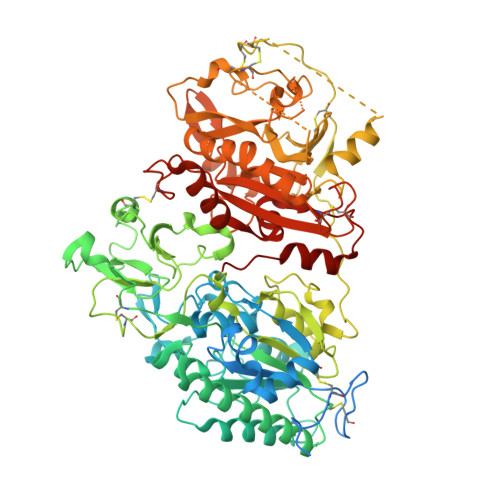
Crystal structure of Enpp1, an extracellular glycoprotein involved in bone mineralization and insulin signaling.
Kato, K., Nishimasu, H., Okudaira, S., Mihara, E., Ishitani, R., Takagi, J., Aoki, J., Nureki, O.(2012) Proc Natl Acad Sci U S A 109: 16876-16881
- PubMed: 23027977
- DOI: https://doi.org/10.1073/pnas.1208017109
- Primary Citation of Related Structures:
4GTW, 4GTX, 4GTY, 4GTZ - PubMed Abstract:
Enpp1 is a membrane-bound glycoprotein that regulates bone mineralization by hydrolyzing extracellular nucleotide triphosphates to produce pyrophosphate. Enpp1 dysfunction causes human diseases characterized by ectopic calcification. Enpp1 also inhibits insulin signaling, and an Enpp1 polymorphism is associated with insulin resistance. However, the precise mechanism by which Enpp1 functions in these cellular processes remains elusive. Here, we report the crystal structures of the extracellular region of mouse Enpp1 in complex with four different nucleotide monophosphates, at resolutions of 2.7-3.2 Å. The nucleotides are accommodated in a pocket formed by an insertion loop in the catalytic domain, explaining the preference of Enpp1 for an ATP substrate. Structural mapping of disease-associated mutations indicated the functional importance of the interdomain interactions. A structural comparison of Enpp1 with Enpp2, a lysophospholipase D, revealed marked differences in the domain arrangements and active-site architectures. Notably, the Enpp1 mutant lacking the insertion loop lost the nucleotide-hydrolyzing activity but instead gained the lysophospholipid-hydrolyzing activity of Enpp2. Our findings provide structural insights into how the Enpp family proteins evolved to exert their diverse cellular functions.
Organizational Affiliation:
Department of Biophysics and Biochemistry, Graduate School of Science, University of Tokyo, Bunkyo-ku, Tokyo 113-0032, Japan.