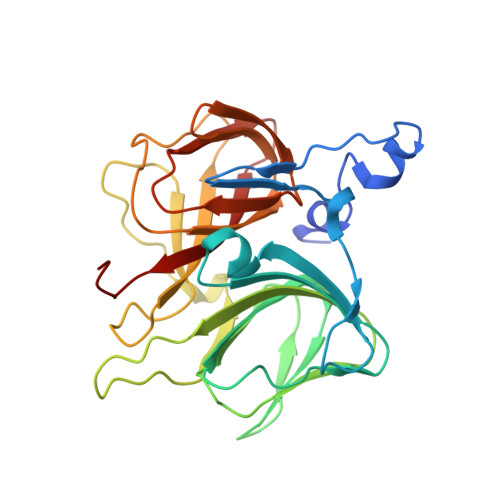
Structural and functional insights into (S)-ureidoglycine aminohydrolase, key enzyme of purine catabolism in Arabidopsis thaliana
Shin, I., Percudani, R., Rhee, S.(2012) J Biol Chem 287: 18796-18805
- PubMed: 22493446
- DOI: https://doi.org/10.1074/jbc.M111.331819
- Primary Citation of Related Structures:
4E2Q, 4E2S - PubMed Abstract:
The ureide pathway has recently been identified as the metabolic route of purine catabolism in plants and some bacteria. In this pathway, uric acid, which is a major product of the early stage of purine catabolism, is degraded into glyoxylate and ammonia via stepwise reactions of seven different enzymes. Therefore, the pathway has a possible physiological role in mobilization of purine ring nitrogen for further assimilation. (S)-Ureidoglycine aminohydrolase enzyme converts (S)-ureidoglycine into (S)-ureidoglycolate and ammonia, providing the final substrate to the pathway. Here, we report a structural and functional analysis of this enzyme from Arabidopsis thaliana (AtUGlyAH). The crystal structure of AtUGlyAH in the ligand-free form shows a monomer structure in the bicupin fold of the β-barrel and an octameric functional unit as well as a Mn(2+) ion binding site. The structure of AtUGlyAH in complex with (S)-ureidoglycine revealed that the Mn(2+) ion acts as a molecular anchor to bind (S)-ureidoglycine, and its binding mode dictates the enantioselectivity of the reaction. Further kinetic analysis characterized the functional roles of the active site residues, including the Mn(2+) ion binding site and residues in the vicinity of (S)-ureidoglycine. These analyses provide molecular insights into the structure of the enzyme and its possible catalytic mechanism.
Organizational Affiliation:
Department of Agricultural Biotechnology, Seoul National University, Seoul, Korea.