Evaluation of Synthetic FK506 Analogues as Ligands for the FK506-Binding Proteins 51 and 52.
Gopalakrishnan, R., Kozany, C., Gaali, S., Kress, C., Hoogeland, B., Bracher, A., Hausch, F.(2012) J Med Chem 55: 4114-4122
- PubMed: 22455444
- DOI: https://doi.org/10.1021/jm201746x
- Primary Citation of Related Structures:
4DRK, 4DRM, 4DRN, 4DRO, 4DRP - PubMed Abstract:
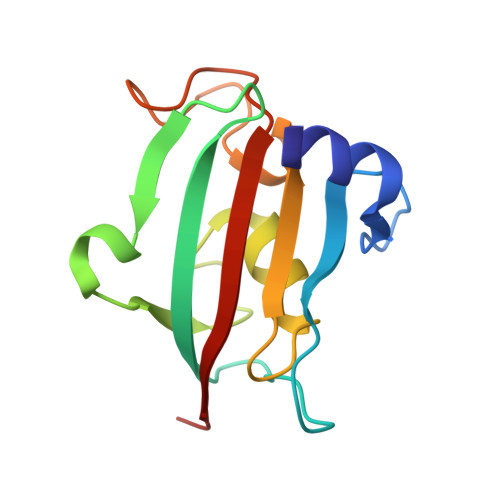
The FK506-binding proteins (FKBP) 51 and 52 are cochaperones that modulate the signal transduction of steroid hormone receptors. Both proteins have been implicated in prostate cancer. Furthermore, single nucleotide polymorphisms in the gene encoding FKBP51 have been associated with a variety of psychiatric disorders. Rapamycin and FK506 are two macrocyclic natural products that bind to these proteins indiscriminately but with nanomolar affinity. We here report the cocrystal structure of FKBP51 with a simplified α-ketoamide analogue derived from FK506 and the first structure-activity relationship analysis for FKBP51 and FKBP52 based on this compound. In particular, the tert-pentyl group of this ligand was systematically replaced by a cyclohexyl ring system, which more closely resembles the pyranose ring in the high-affinity ligands rapamycin and FK506. The interaction with FKBPs was found to be surprisingly tolerant to the stereochemistry of the attached cyclohexyl substituents. The molecular basis for this tolerance was elucidated by X-ray cocrystallography.
Organizational Affiliation:
Max Planck Institute of Psychiatry, Kraepelinstrasse 2, 80804 Munich, Germany.