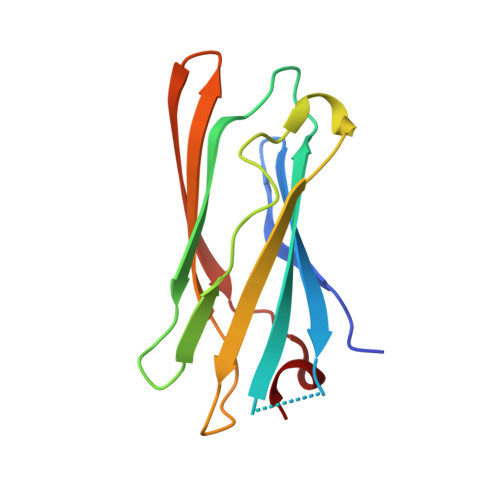
Computational Design of a Symmetric Homodimer Using Beta-Strand Assembly.
Stranges, P.B., Machius, M., Miley, M.J., Tripathy, A., Kuhlman, B.(2011) Proc Natl Acad Sci U S A 108: 20562
- PubMed: 22143762
- DOI: https://doi.org/10.1073/pnas.1115124108
- Primary Citation of Related Structures:
3ZY7 - PubMed Abstract:
Computational design of novel protein-protein interfaces is a test of our understanding of protein interactions and has the potential to allow modification of cellular physiology. Methods for designing high-affinity interactions that adopt a predetermined binding mode have proved elusive, suggesting the need for new strategies that simplify the design process. A solvent-exposed backbone on a β-strand is thought of as "sticky" and β-strand pairing stabilizes many naturally occurring protein complexes. Here, we computationally redesign a monomeric protein to form a symmetric homodimer by pairing exposed β-strands to form an intermolecular β-sheet. A crystal structure of the designed complex closely matches the computational model (rmsd = 1.0 Å). This work demonstrates that β-strand pairing can be used to computationally design new interactions with high accuracy.
Organizational Affiliation:
Department of Biochemistry and Biophysics, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA.