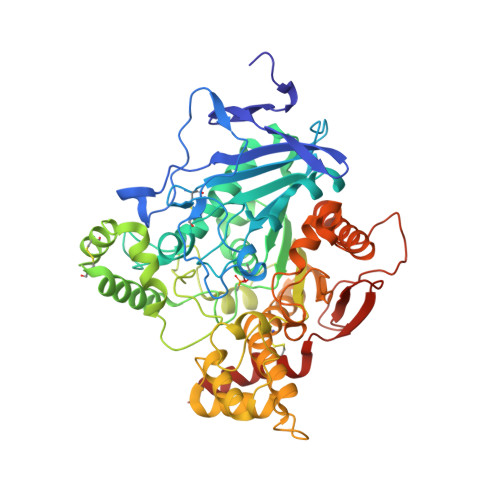
Catalytic-Site Conformational Equilibrium in Nerve-Agent Adducts of Acetylcholinesterase; Possible Implications for the Hi-6 Antidote Substrate Specificity.
Artursson, E., Andersson, P.O., Akfur, C., Linusson, A., Borjegren, S., Ekstrom, F.(2013) Biochem Pharmacol 85: 1389
- PubMed: 23376121
- DOI: https://doi.org/10.1016/j.bcp.2013.01.016
- Primary Citation of Related Structures:
3ZLT, 3ZLU, 3ZLV - PubMed Abstract:
Nerve agents such as tabun, cyclosarin and Russian VX inhibit the essential enzyme acetylcholinesterase (AChE) by organophosphorylating the catalytic serine residue. Nucleophiles, such as oximes, are used as antidotes as they can reactivate and restore the function of the inhibited enzyme. The oxime HI-6 shows a notably low activity on tabun adducts but can effectively reactivate adducts of cyclosarin and Russian VX. To examine the structural basis for the pronounced substrate specificity of HI-6, we determined the binary crystal structures of Mus musculus AChE (mAChE) conjugated by cyclosarin and Russian VX and found a conformational mobility of the side chains of Phe338 and His447. The interaction between HI-6 and tabun-adducts of AChE were subsequently investigated using a combination of time resolved fluorescence spectroscopy and X-ray crystallography. Our findings show that HI-6 binds to tabun inhibited Homo sapiens AChE (hAChE) with an IC50 value of 300μM and suggest that the reactive nucleophilic moiety of HI-6 is excluded from the phosphorus atom of tabun. We propose that a conformational mobility of the side-chains of Phe338 and His447 is a common feature in nerve-agent adducts of AChE. We also suggest that the conformational mobility allow HI-6 to reactivate conjugates of cyclosarin and Russian VX while a reduced mobility in tabun conjugated AChE results in steric hindrance that prevents efficient reactivation.
Organizational Affiliation:
Swedish Defence Research Agency, CBRN, Defence and Security, Umeå, Sweden.