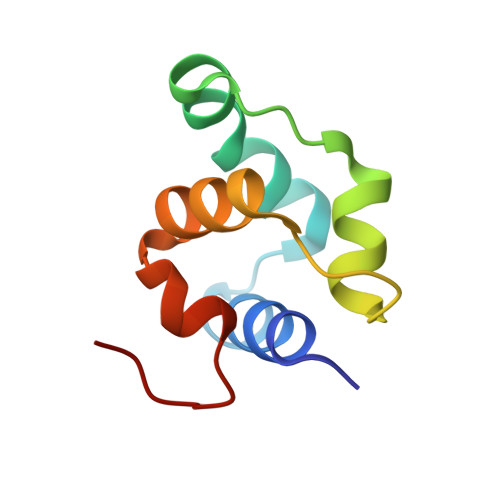
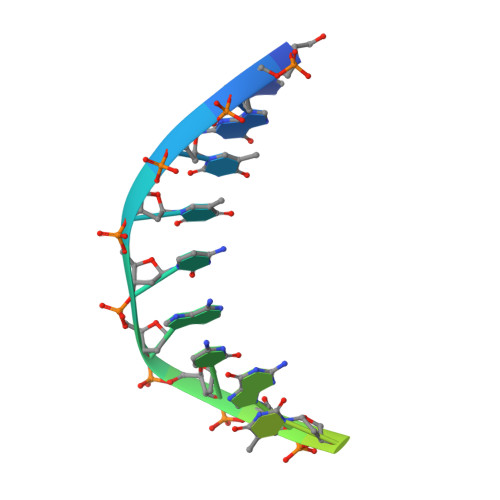
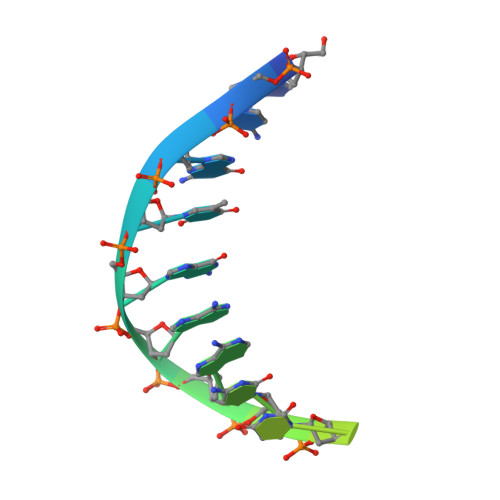
Binding of the N-Terminal Domain of the Lactococcal Bacteriophage Tp901-1 Ci Repressor to its Target DNA: A Crystallography, Small Angle Scattering, and Nuclear Magnetic Resonance Study.
Frandsen, K.H., Rasmussen, K.K., Jensen, M.R., Hammer, K., Pedersen, M., Poulsen, J.N., Arleth, L., Lo Leggio, L.(2013) Biochemistry 52: 6892
- PubMed: 24047404
- DOI: https://doi.org/10.1021/bi400439y
- Primary Citation of Related Structures:
3ZHI, 3ZHM - PubMed Abstract:
In most temperate bacteriophages, regulation of the choice of lysogenic or lytic life cycle is controlled by a CI repressor protein. Inhibition of transcription is dependent on a helix-turn-helix motif, often located in the N-terminal domain (NTD), which binds to specific DNA sequences (operator sites). Here the crystal structure of the NTD of the CI repressor from phage TP901-1 has been determined at 1.6 Å resolution, and at 2.6 Å resolution in complex with a 9 bp double-stranded DNA fragment that constitutes a half-site of the OL operator. This N-terminal construct, comprising residues 2-74 of the CI repressor, is monomeric in solution as shown by nuclear magnetic resonance (NMR), small angle X-ray scattering, and gel filtration and is monomeric in the crystal structures. The binding interface between the NTD and the half-site in the crystal is very similar to the interface that can be mapped by NMR in solution with a full palindromic site. The interactions seen in the complexes (in the crystal and in solution) explain the observed affinity for the OR site that is lower than that for the OL site and the specificity for the recognized DNA sequence in comparison to that for other repressors. Compared with many well-studied phage repressor systems, the NTD from TP901-1 CI has a longer extended scaffolding helix that, interestingly, is strongly conserved in putative repressors of Gram-positive pathogens. On the basis of sequence comparisons, we suggest that these bacteria also possess repressor/antirepressor systems similar to that found in phage TP901-1.
Organizational Affiliation:
Department of Chemistry, University of Copenhagen , Universitetsparken 5, DK-2100 Copenhagen Ø, Denmark.