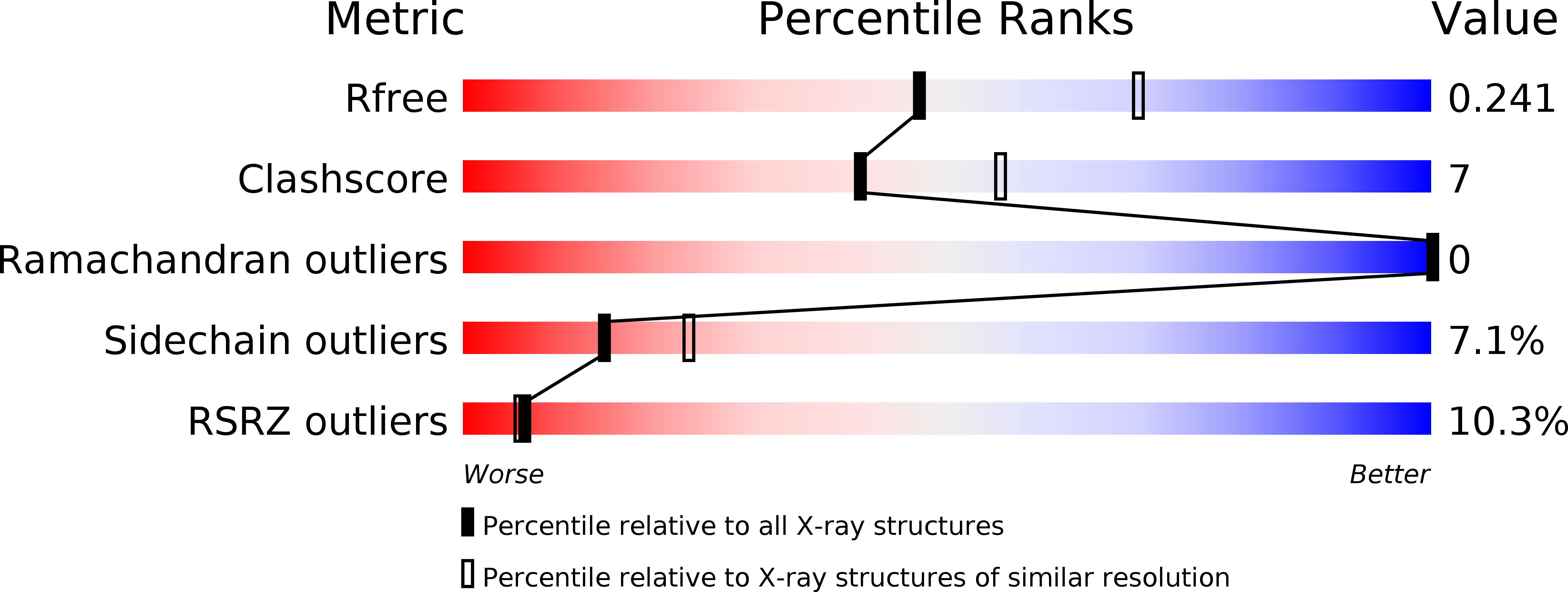
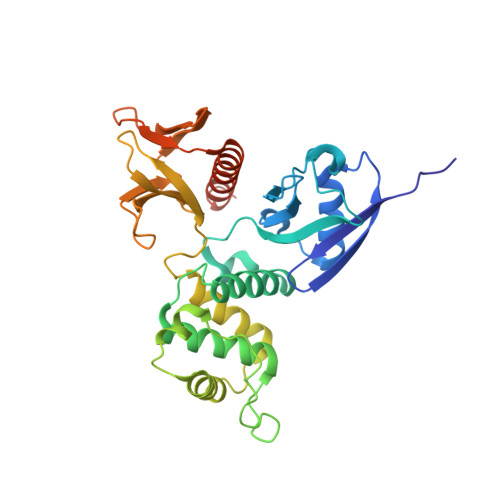
MT1-MMP recognition by ERM proteins and its implication in CD44 shedding
Terawaki, S., Kitano, K., Aoyama, M., Mori, T., Hakoshima, T.(2015) Genes Cells 20: 847-859
- PubMed: 26289026
- DOI: https://doi.org/10.1111/gtc.12276
- Primary Citation of Related Structures:
3X23 - PubMed Abstract:
Membrane type 1-matrix metalloproteinase (MT1-MMP) is a key enzyme involved in tumor cell invasion by shedding their cell-surface receptor CD44 anchored with F-actin through ezrin/radixin/moesin (ERM) proteins. We found the cytoplasmic tail of MT1-MMP directly binds the FERM domain of radixin, suggesting F-actin-based recruitment of MT1-MMP to CD44 for invasion. Our crystal structure shows that the central region of the MT1-MMP cytoplasmic tail binds subdomain A of the FERM domain, and makes an antiparallel β-β interaction with β2A-strand. This binding mode is distinct from the previously determined binding mode of CD44 to subdomain C. We showed that radixin simultaneously binds both MT1-MMP and CD44, indicating ERM protein-mediated colocalization of MT1-MMP and its substrate CD44 and anchoring to F-actin. Our study implies that ERM proteins contribute toward accelerated CD44 shedding by MT1-MMP through ERM protein-mediated interactions between their cytoplasmic tails.
Organizational Affiliation:
Structural Biology Laboratory, Nara Institute of Science and Technology, 8916-5 Takayama, Ikoma, Nara, 630-0192, Japan.