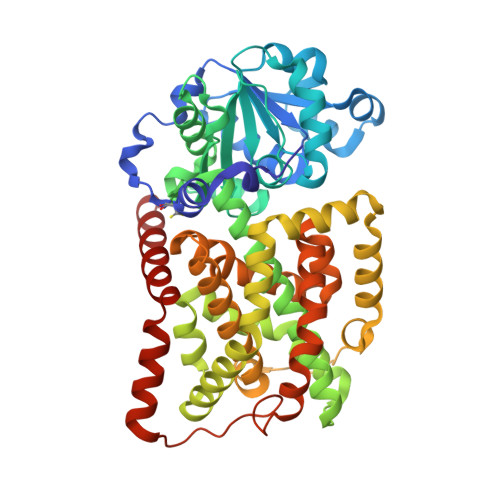
Bacterial and plant ketol-acid reductoisomerases have different mechanisms of induced fit during the catalytic cycle
Wong, S.H., Lonhienne, T.G.A., Winzor, D.J., Schenk, G., Guddat, L.W.(2012) J Mol Biol 424: 168-179
- PubMed: 23036858
- DOI: https://doi.org/10.1016/j.jmb.2012.09.018
- Primary Citation of Related Structures:
3ULK - PubMed Abstract:
Ketol-acid reductoisomerase (KARI) is the second enzyme in the branched-chain amino acid biosynthesis pathway, which is found in plants, fungi and bacteria but not in animals. This difference in metabolism between animals and microorganisms makes KARI an attractive target for the development of antimicrobial agents. Herein we report the crystal structure of Escherichia coli KARI in complex with Mg(2+) and NADPH at 2.3Å resolution. Ultracentrifugation studies confirm that the enzyme exists as a tetramer in solution, and isothermal titration calorimetry shows that the binding of Mg(2+) increases structural disorder while the binding of NADPH increases the structural rigidity of the enzyme. Comparison of the structure of the E. coli KARI-Mg(2+)-NADPH complex with that of enzyme in the absence of cofactors shows that the binding of Mg(2+) and NADPH opens the interface between the N- and C-domains, thereby allowing access for the substrates to bind: the existence of only a small opening between the domains in the crystal structure of the unliganded enzyme signifies restricted access to the active site. This observation contrasts with that in the plant enzyme, where the N-domain can rotate freely with respect to the C-domain until the binding of Mg(2+) and/or NADPH stabilizes the relative positions of these domains. Support is thereby provided for the idea that plant and bacterial KARIs have evolved different mechanisms of induced fit to prepare the active site for catalysis.
Organizational Affiliation:
School of Chemistry and Molecular Biosciences, The University of Queensland, Brisbane, Queensland 4072, Australia. s.wong3@uq.edu.au