Structural characterization of beta-ketoacyl ACP synthase I bound to platencin and fragment screening molecules at two substrate binding sites.
Patterson, E.I., Nanson, J.D., Abendroth, J., Bryan, C., Sankaran, B., Myler, P.J., Forwood, J.K.(2020) Proteins 88: 47-56
- PubMed: 31237717
- DOI: https://doi.org/10.1002/prot.25765
- Primary Citation of Related Structures:
3LRF, 3MQD, 3U0E, 3U0F, 4JV3 - PubMed Abstract:
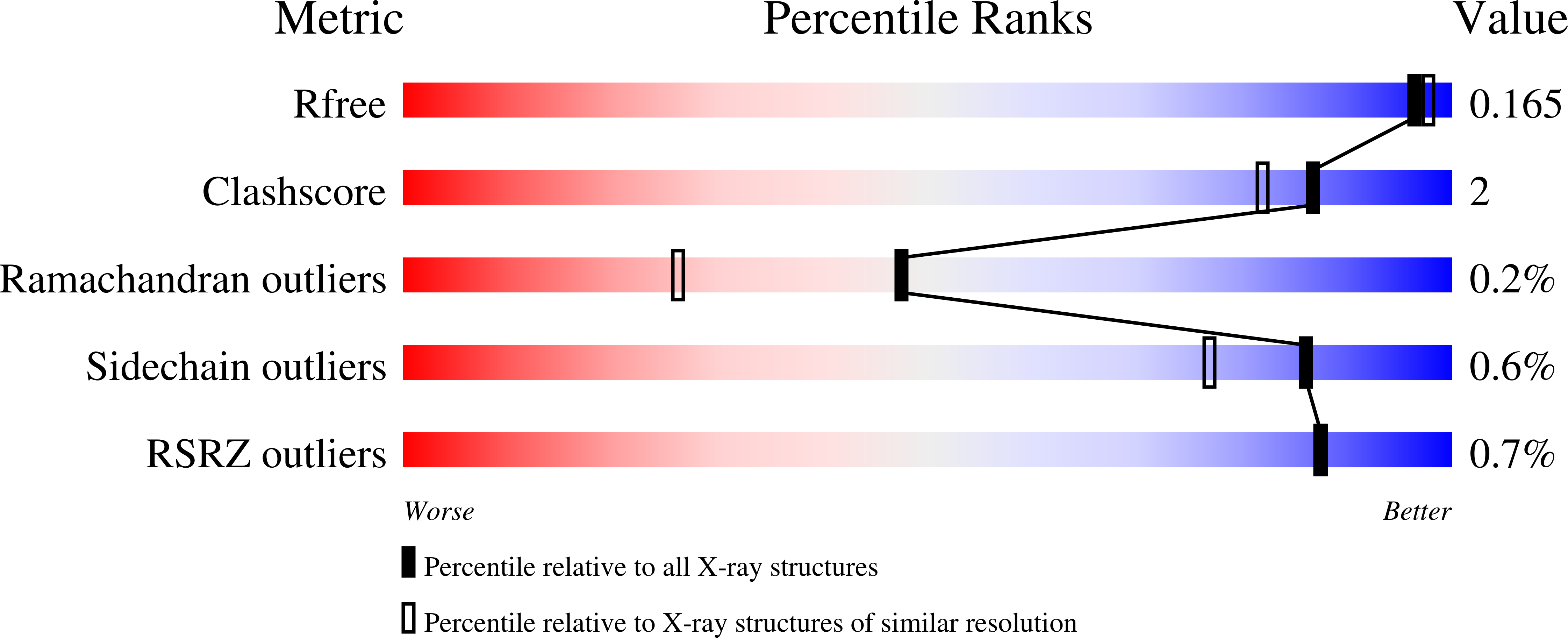
The bacterial fatty acid pathway is essential for membrane synthesis and a range of other metabolic and cellular functions. The β-ketoacyl-ACP synthases carry out the initial elongation reaction of this pathway, utilizing acetyl-CoA as a primer to elongate malonyl-ACP by two carbons, and subsequent elongation of the fatty acyl-ACP substrate by two carbons. Here we describe the structures of the β-ketoacyl-ACP synthase I from Brucella melitensis in complex with platencin, 7-hydroxycoumarin, and (5-thiophen-2-ylisoxazol-3-yl)methanol. The enzyme is a dimer and based on structural and sequence conservation, harbors the same active site configuration as other β-ketoacyl-ACP synthases. The platencin binding site overlaps with the fatty acyl compound supplied by ACP, while 7-hydroxyl-coumarin and (5-thiophen-2-ylisoxazol-3-yl)methanol bind at the secondary fatty acyl binding site. These high-resolution structures, ranging between 1.25 and 1.70 å resolution, provide a basis for in silico inhibitor screening and optimization, and can aid in rational drug design by revealing the high-resolution binding interfaces of molecules at the malonyl-ACP and acyl-ACP active sites.
Organizational Affiliation:
Department of Pathology, University of Texas Medical Branch, Galveston, Texas.