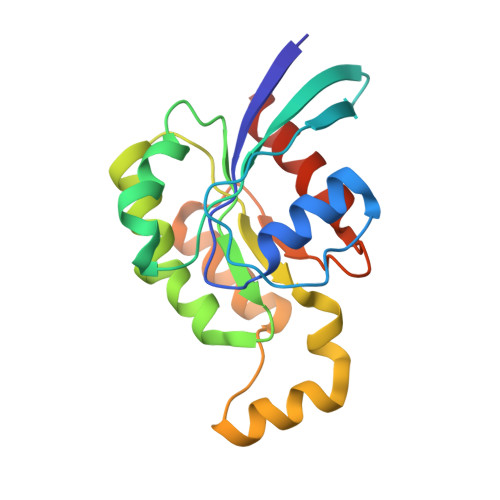
A Dual Binding Mode for RhoGTPases in Plexin Signalling.
Bell, C.H., Aricescu, A.R., Jones, E.Y., Siebold, C.(2011) PLoS Biol 9: e1001134-e1001134
- PubMed: 21912513
- DOI: https://doi.org/10.1371/journal.pbio.1001134
- Primary Citation of Related Structures:
3SU8, 3SUA - PubMed Abstract:
Plexins are cell surface receptors for the semaphorin family of cell guidance cues. The cytoplasmic region comprises a Ras GTPase-activating protein (GAP) domain and a RhoGTPase binding domain. Concomitant binding of extracellular semaphorin and intracellular RhoGTPase triggers GAP activity and signal transduction. The mechanism of this intricate regulation remains elusive. We present two crystal structures of the human Plexin-B1 cytoplasmic region in complex with a constitutively active RhoGTPase, Rac1. The structure of truncated Plexin-B1-Rac1 complex provides no mechanism for coupling RhoGTPase and Ras binding sites. On inclusion of the juxtamembrane helix, a trimeric structure of Plexin-B1-Rac1 complexes is stabilised by a second, novel, RhoGTPase binding site adjacent to the Ras site. Site-directed mutagenesis combined with cellular and biophysical assays demonstrate that this new binding site is essential for signalling. Our findings are consistent with a model in which extracellular and intracellular plexin clustering events combine into a single signalling output.
Organizational Affiliation:
Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, United Kingdom.