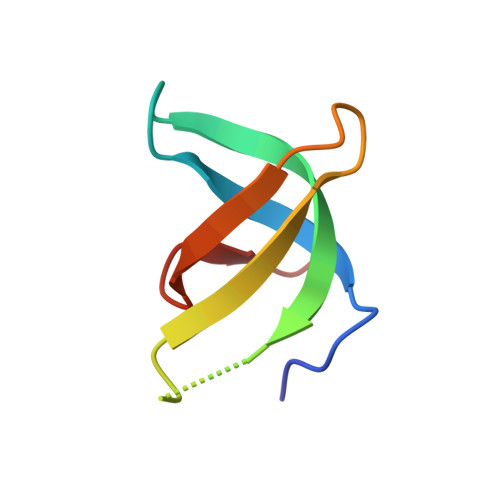
Crystal structure of TDRD3 and methyl-arginine binding characterization of TDRD3, SMN and SPF30
Liu, K., Guo, Y.H., Liu, H.P., Bian, C.B., Lam, R., Liu, Y., Mackenzie, F., Rojas, L.A., Reinberg, D., Bedford, M.T., Xu, R.M., Min, J.R.(2012) PLoS One 7: e30375-e30375
- PubMed: 22363433
- DOI: https://doi.org/10.1371/journal.pone.0030375
- Primary Citation of Related Structures:
3PMT, 3S6W - PubMed Abstract:
SMN (Survival motor neuron protein) was characterized as a dimethyl-arginine binding protein over ten years ago. TDRD3 (Tudor domain-containing protein 3) and SPF30 (Splicing factor 30 kDa) were found to bind to various methyl-arginine proteins including Sm proteins as well later on. Recently, TDRD3 was shown to be a transcriptional coactivator, and its transcriptional activity is dependent on its ability to bind arginine-methylated histone marks. In this study, we systematically characterized the binding specificity and affinity of the Tudor domains of these three proteins quantitatively. Our results show that TDRD3 preferentially recognizes asymmetrical dimethylated arginine mark, and SMN is a very promiscuous effector molecule, which recognizes different arginine containing sequence motifs and preferentially binds symmetrical dimethylated arginine. SPF30 is the weakest methyl-arginine binder, which only binds the GAR motif sequences in our library. In addition, we also reported high-resolution crystal structures of the Tudor domain of TDRD3 in complex with two small molecules, which occupy the aromatic cage of TDRD3.
Organizational Affiliation:
Hubei Key Laboratory of Genetic Regulation and Integrative Biology, College of Life Science, Huazhong Normal University, Wuhan, People's Republic of China.