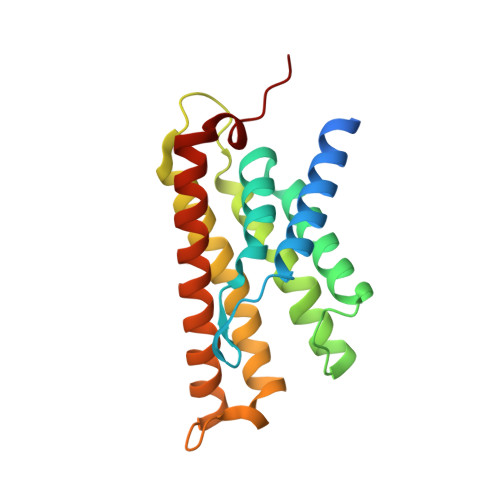
The structural basis of modularity in ECF-type ABC transporters.
Erkens, G.B., Berntsson, R.P., Fulyani, F., Majsnerowska, M., Ter Beek, J., Poolman, B., Slotboom, D.J.(2011) Nat Struct Mol Biol 18: 755-760
- PubMed: 21706007
- DOI: https://doi.org/10.1038/nsmb.2073
- Primary Citation of Related Structures:
3RLB - PubMed Abstract:
Energy coupling factor (ECF) transporters are used for the uptake of vitamins in Prokarya. They consist of an integral membrane protein that confers substrate specificity (the S-component) and an energizing module that is related to ATP-binding cassette (ABC) transporters. S-components for different substrates often do not share detectable sequence similarity but interact with the same energizing module. Here we present the crystal structure of the thiamine-specific S-component ThiT from Lactococcus lactis at 2.0 Å. Extensive protein-substrate interactions explain its high binding affinity for thiamine (K(d) ~10(-10) M). ThiT has a fold similar to that of the riboflavin-specific S-component RibU, with which it shares only 14% sequence identity. Two alanines in a conserved motif (AxxxA) located on the membrane-embedded surface of the S-components mediate the interaction with the energizing module. Based on these findings, we propose a general transport mechanism for ECF transporters.
Organizational Affiliation:
University of Groningen, Groningen Biomolecular Science and Biotechnology Institute, Groningen, The Netherlands.