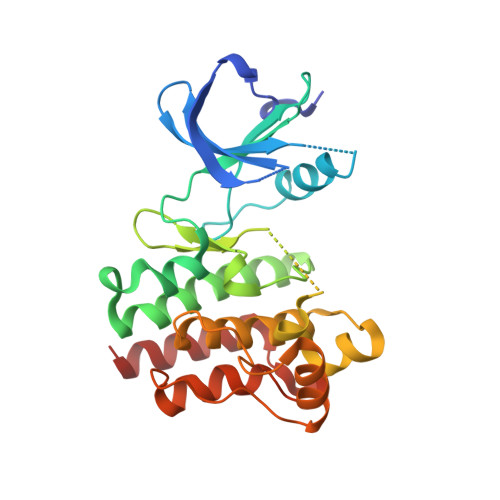
Insights into the conformational flexibility of Bruton's tyrosine kinase from multiple ligand complex structures.
Kuglstatter, A., Wong, A., Tsing, S., Lee, S.W., Lou, Y., Villasenor, A.G., Bradshaw, J.M., Shaw, D., Barnett, J.W., Browner, M.F.(2011) Protein Sci 20: 428-436
- PubMed: 21280133
- DOI: https://doi.org/10.1002/pro.575
- Primary Citation of Related Structures:
3PIX, 3PIY, 3PIZ, 3PJ1, 3PJ2, 3PJ3 - PubMed Abstract:
Bruton's tyrosine kinase (BTK) plays a key role in B cell receptor signaling and is considered a promising drug target for lymphoma and inflammatory diseases. We have determined the X-ray crystal structures of BTK kinase domain in complex with six inhibitors from distinct chemical classes. Five different BTK protein conformations are stabilized by the bound inhibitors, providing insights into the structural flexibility of the Gly-rich loop, helix C, the DFG sequence, and activation loop. The conformational changes occur independent of activation loop phosphorylation and do not correlate with the structurally unchanged WEI motif in the Src homology 2-kinase domain linker. Two novel activation loop conformations and an atypical DFG conformation are observed representing unique inactive states of BTK. Two regions within the activation loop are shown to structurally transform between 3(10)- and α-helices, one of which collapses into the adenosine-5'-triphosphate binding pocket. The first crystal structure of a Tec kinase family member in the pharmacologically important DFG-out conformation and bound to a type II kinase inhibitor is described. The different protein conformations observed provide insights into the structural flexibility of BTK, the molecular basis of its regulation, and the structure-based design of specific inhibitors.
Organizational Affiliation:
Roche Palo Alto, 3431 Hillview Avenue, Palo Alto, California 94304, USA. andreas.kuglstatter@roche.com