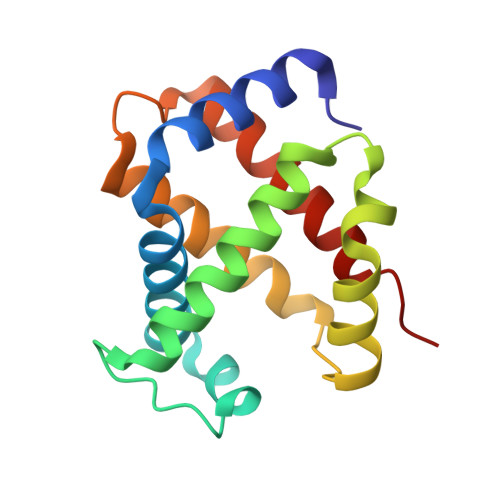
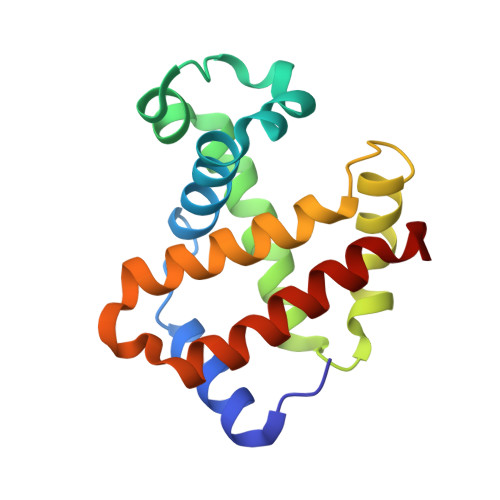
Blocking the gate to ligand entry in human hemoglobin.
Birukou, I., Soman, J., Olson, J.S.(2011) J Biol Chem 286: 10515-10529
- PubMed: 21193395
- DOI: https://doi.org/10.1074/jbc.M110.176271
- Primary Citation of Related Structures:
3NL7, 3NML, 3NMM, 3OGB - PubMed Abstract:
His(E7) to Trp replacements in HbA lead to markedly biphasic bimolecular CO rebinding after laser photolysis. For isolated mutant subunits, the fraction of fast phase increases with increasing [CO], suggesting a competition between binding to an open conformation with an empty E7 channel and relaxation to blocked or closed, slowly reacting states. The rate of conformational relaxation of the open state is ∼18,000 s(-1) in α subunits and ∼10-fold faster in β subunits, ∼175,000 s(-1). Crystal structures were determined for tetrameric α(WT)β(Trp-63) HbCO, α(Trp-58)β(WT) deoxyHb, and Trp-64 deoxy- and CO-Mb as controls. In Trp-63(E7) βCO, the indole side chain is located in the solvent interface, blocking entry into the E7 channel. Similar blocked Trp-64(E7) conformations are observed in the mutant Mb crystal structures. In Trp-58(E7) deoxy-α subunits, the indole side chain fills both the channel and the distal pocket, forming a completely closed state. The bimolecular rate constant for CO binding, k'(CO), to the open conformations of both mutant Hb subunits is ∼80-90 μm(-1) s(-1), whereas k'(CO) for the completely closed states is 1000-fold slower, ∼0.08 μm(-1) s(-1). A transient intermediate with k'(CO) ≈ 0.7 μm(-1) s(-1) is observed after photolysis of Trp-63(E7) βCO subunits and indicates that the indole ring blocks the entrance to the E7 channel, as observed in the crystal structures of Trp(E7) deoxyMb and βCO subunits. Thus, either blocking or completely filling the E7 channel dramatically slows bimolecular binding, providing strong evidence that the E7 channel is the major pathway (≥90%) for ligand entry in human hemoglobin.
Organizational Affiliation:
Department of Biochemistry and Cell Biology and the W. M. Keck Center for Computational Biology, Rice University, Houston, Texas 77005, USA.