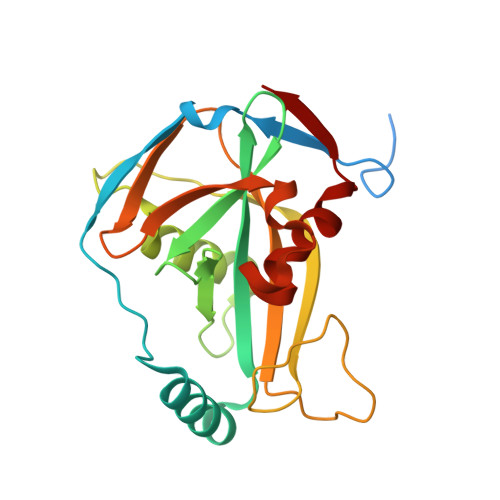
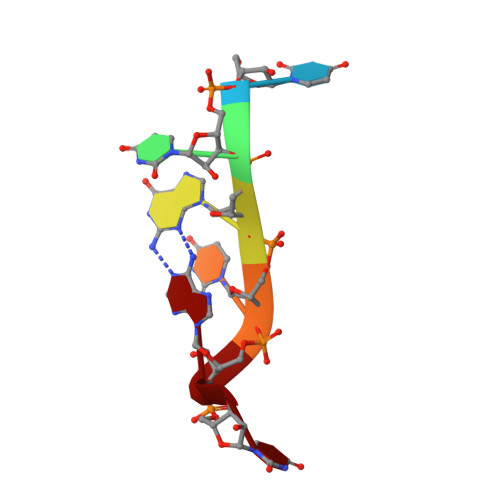
Structural basis of UGUA recognition by the Nudix protein CFI(m)25 and implications for a regulatory role in mRNA 3' processing.
Yang, Q., Gilmartin, G.M., Doublie, S.(2010) Proc Natl Acad Sci U S A 107: 10062-10067
- PubMed: 20479262
- DOI: https://doi.org/10.1073/pnas.1000848107
- Primary Citation of Related Structures:
3MDG, 3MDI - PubMed Abstract:
Human Cleavage Factor Im (CFI(m)) is an essential component of the pre-mRNA 3' processing complex that functions in the regulation of poly(A) site selection through the recognition of UGUA sequences upstream of the poly(A) site. Although the highly conserved 25 kDa subunit (CFI(m)25) of the CFI(m) complex possesses a characteristic alpha/beta/alpha Nudix fold, CFI(m)25 has no detectable hydrolase activity. Here we report the crystal structures of the human CFI(m)25 homodimer in complex with UGUAAA and UUGUAU RNA sequences. CFI(m)25 is the first Nudix protein to be reported to bind RNA in a sequence-specific manner. The UGUA sequence contributes to binding specificity through an intramolecular G:A Watson-Crick/sugar-edge base interaction, an unusual pairing previously found to be involved in the binding specificity of the SAM-III riboswitch. The structures, together with mutational data, suggest a novel mechanism for the simultaneous sequence-specific recognition of two UGUA elements within the pre-mRNA. Furthermore, the mutually exclusive binding of RNA and the signaling molecule Ap(4)A (diadenosine tetraphosphate) by CFI(m)25 suggests a potential role for small molecules in the regulation of mRNA 3' processing.
Organizational Affiliation:
Department of Microbiology and Molecular Genetics, Stafford Hall, University of Vermont, Burlington, VT 05405, USA.