Crystal structures of aprotinin and its complex with sucrose octasulfate reveal multiple modes of interactions with implications for heparin binding
Yang, I.S., Kim, T.G., Park, B.S., Cho, K.J., Lee, J.-H., Park, Y., Kim, K.H.(2010) Biochem Biophys Res Commun 397: 429-435
- PubMed: 20529698
- DOI: https://doi.org/10.1016/j.bbrc.2010.05.113
- Primary Citation of Related Structures:
3LDI, 3LDJ, 3LDM - PubMed Abstract:
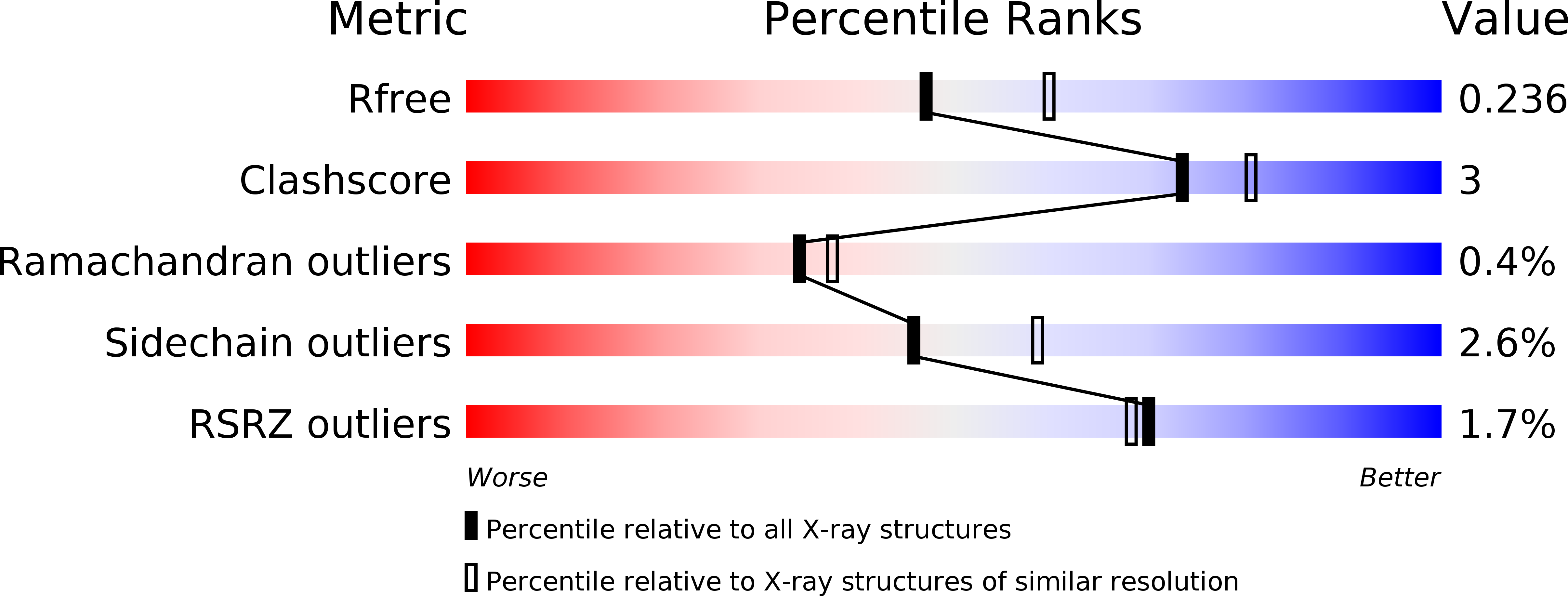
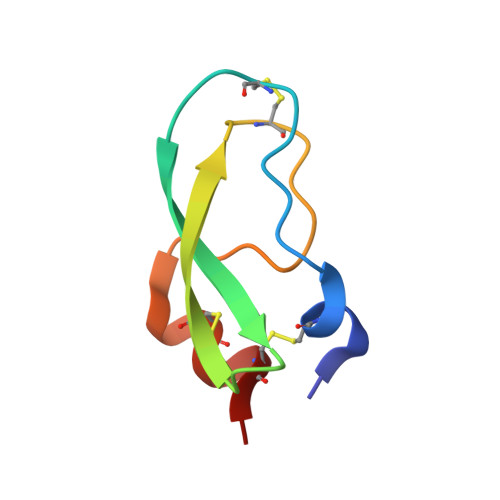
The crystal structures of aprotinin and its complex with sucrose octasulfate (SOS), a polysulfated heparin analog, were determined at 1.7-2.6A resolutions. Aprotinin is monomeric in solution, which associates into a decamer at high salt concentrations. Sulfate ions serve to neutralize the basic amino acid residues of aprotinin to stabilize the decameric aprotinin. Whereas SOS interacts with heparin binding proteins at 1:1 molar ratio, SOS was surprisingly found to induce strong agglutination of aprotinins. Five molecules of aprotinin interact with one molecule of the sulfated sugar, which is stabilized by electrostatic interactions between the positively charged residues of aprotinin and sulfate groups of SOS. The multiple binding modes of SOS with five individual aprotinin molecules may represent the diverse patterns of potential heparin binding to aprotinin, reflecting the interactions of densely packed protein molecules along the heparin polymer.
Organizational Affiliation:
Department of Biotechnology & Bioinformatics, Korea University, Chungnam 339-700, Republic of Korea.