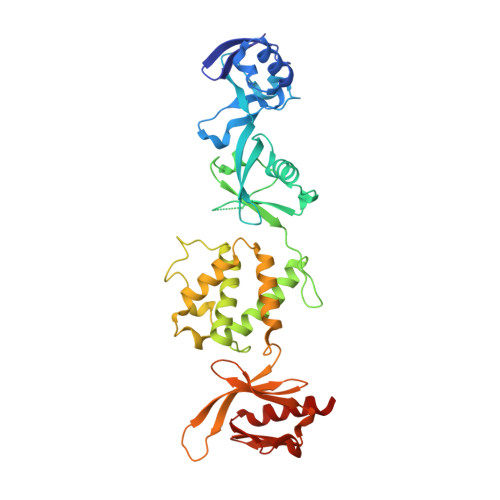
The Structure of the talin head reveals a novel extended conformation of the FERM domain
Elliott, P.R., Goult, B.T., Kopp, P.M., Bate, N., Grossmann, J.G., Roberts, G.C.K., Critchley, D.R., Barsukov, I.L.(2010) Structure 18: 1289-1299
- PubMed: 20947018
- DOI: https://doi.org/10.1016/j.str.2010.07.011
- Primary Citation of Related Structures:
3IVF - PubMed Abstract:
FERM domains are found in a diverse superfamily of signaling and adaptor proteins at membrane interfaces. They typically consist of three separately folded domains (F1, F2, F3) in a compact cloverleaf structure. The crystal structure of the N-terminal head of the integrin-associated cytoskeletal protein talin reported here reveals a novel FERM domain with a linear domain arrangement, plus an additional domain F0 packed against F1. While F3 binds β-integrin tails, basic residues in F1 and F2 are required for membrane association and for integrin activation. We show that these same residues are also required for cell spreading and focal adhesion assembly in cells. We suggest that the extended conformation of the talin head allows simultaneous binding to integrins via F3 and to PtdIns(4,5)P2-enriched microdomains via basic residues distributed along one surface of the talin head, and that these multiple interactions are required to stabilize integrins in the activated state.
Organizational Affiliation:
School of Biological Sciences, University of Liverpool, Crown Street, Liverpool, L69 7ZB, UK.