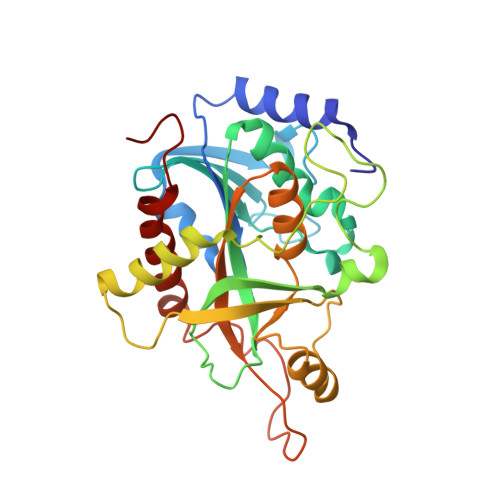
Crystal structure and molecular dynamics studies of human purine nucleoside phosphorylase complexed with 7-deazaguanine.
Caceres, R.A., Timmers, L.F., Pauli, I., Gava, L.M., Ducati, R.G., Basso, L.A., Santos, D.S., de Azevedo, W.F.(2010) J Struct Biol 169: 379-388
- PubMed: 19932753
- DOI: https://doi.org/10.1016/j.jsb.2009.11.010
- Primary Citation of Related Structures:
3INY - PubMed Abstract:
In humans, purine nucleoside phosphorylase (HsPNP) is responsible for degradation of deoxyguanosine, and genetic deficiency of this enzyme leads to profound T-cell mediated immunosuppression. HsPNP is a target for inhibitor development aiming at T-cell immune response modulation. Here we report the crystal structure of HsPNP in complex with 7-deazaguanine (HsPNP:7DG) at 2.75 A. Molecular dynamics simulations were employed to assess the structural features of HsPNP in both free form and in complex with 7DG. Our results show that some regions, responsible for entrance and exit of substrate, present a conformational variability, which is dissected by dynamics simulation analysis. Enzymatic assays were also carried out and revealed that 7-deazaguanine presents a lower inhibitory activity against HsPNP (K(i)=200 microM). The present structure may be employed in both structure-based design of PNP inhibitors and in development of specific empirical scoring functions.
Organizational Affiliation:
Faculdade de Biociências, Instituto Nacional de Ciência e Tecnologia em Tuberculose-CNPq, Laboratório de Bioquímica Estrutural, Pontifícia Universidade Católica do Rio Grande do Sul, PUCRS, Porto Alegre, RS, Brazil.