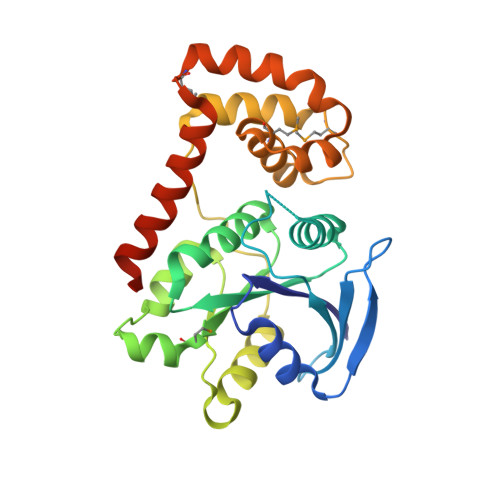
Structural basis of GDP release and gating in G protein coupled Fe2+ transport.
Guilfoyle, A., Maher, M.J., Rapp, M., Clarke, R., Harrop, S., Jormakka, M.(2009) EMBO J 28: 2677-2685
- PubMed: 19629046
- DOI: https://doi.org/10.1038/emboj.2009.208
- Primary Citation of Related Structures:
3HYR, 3HYT - PubMed Abstract:
G proteins are key molecular switches in the regulation of membrane protein function and signal transduction. The prokaryotic membrane protein FeoB is involved in G protein coupled Fe(2+) transport, and is unique in that the G protein is directly tethered to the membrane domain. Here, we report the structure of the soluble domain of FeoB, including the G protein domain, and its assembly into an unexpected trimer. Comparisons between nucleotide free and liganded structures reveal the closed and open state of a central cytoplasmic pore, respectively. In addition, these data provide the first observation of a conformational switch in the nucleotide-binding G5 motif, defining the structural basis for GDP release. From these results, structural parallels are drawn to eukaryotic G protein coupled membrane processes.
Organizational Affiliation:
Centenary Institute, Sydney, New South Wales, Australia.