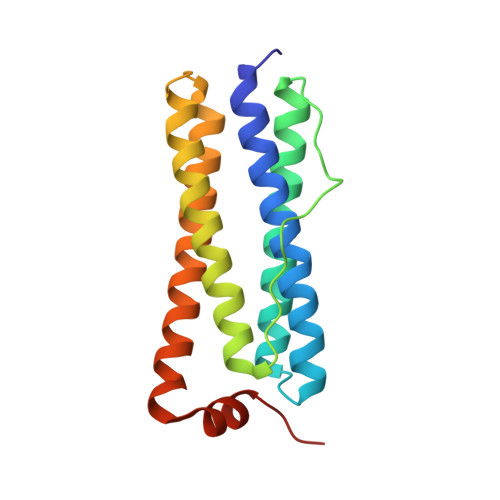
Crystal structure of bacterioferritin from Rhodobacter sphaeroides
Nam, K.H., Xu, Y., Piao, S., Priyadarshi, A., Lee, E.H., Kim, H.-Y., Jeon, Y.H., Ha, N.-C., Hwang, K.Y.(2010) Biochem Biophys Res Commun 391: 990-994
- PubMed: 19968959
- DOI: https://doi.org/10.1016/j.bbrc.2009.12.003
- Primary Citation of Related Structures:
3GVY - PubMed Abstract:
Iron is essential for the survival of organisms, but either excess or deficient levels of iron induce oxidative stress, thereby causing cell damage. As a result, iron regulation is essential for proper cell growth and proliferation in most organisms. Bacterioferritin is a ferritin-like family protein that contains a heme molecule and a ferroxidase site at the di-iron center. This protein plays a primary role in intracellular iron storage for iron homeostasis, as well as in the maintenance of iron in a soluble and non-toxic form. Although several bacterioferritin structures have been determined, no structural studies have successfully elucidated the molecular function of the heme molecule and the ferroxidase center. Here, we report the crystal structure of bacterioferritin from Rhodobacter sphaeroides. This protein exists in a roughly spherical configuration via the assembly of 24 subunits. We describe the oligomeric arrangement, ferroxidase center and heme-binding site based on this structure. The protein contains a single iron-binding configuration in the ferroxidase center, which allows for the release of iron by His130 when the protein is in the intermediate state. The heme molecule in RsBfr is stabilized by shifting of the van der Waals interaction center between the porphyrin of the heme and Trp26. We anticipate that further structural analysis will provide a more complete understanding of the molecular mechanisms of members of the ferritin-like family.
Organizational Affiliation:
Division of Biotechnology, College of Life Sciences & Biotechnology, Korea University, Seoul 136-701, Republic of Korea.