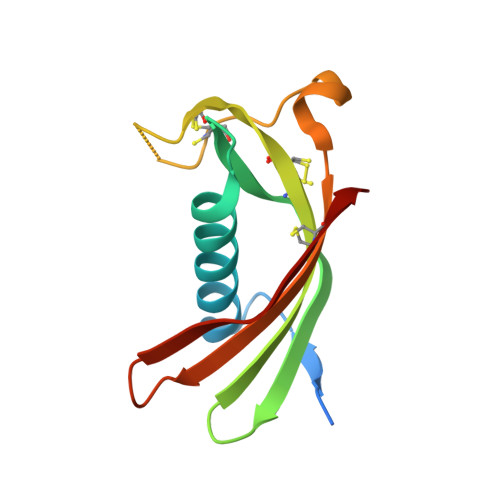
Crystal structure of human cystatin C stabilized against amyloid formation.
Kolodziejczyk, R., Michalska, K., Hernandez-Santoyo, A., Wahlbom, M., Grubb, A., Jaskolski, M.(2010) FEBS J 277: 1726-1737
- PubMed: 20175878
- DOI: https://doi.org/10.1111/j.1742-4658.2010.07596.x
- Primary Citation of Related Structures:
3GAX - PubMed Abstract:
Human cystatin C (HCC) is a family 2 cystatin inhibitor of papain-like (C1) and legumain-related (C13) cysteine proteases. In pathophysiological processes, the nature of which is not understood, HCC is codeposited in the amyloid plaques of Alzheimer's disease or Down's syndrome. The amyloidogenic properties of HCC are greatly increased in a naturally occurring L68Q variant, resulting in fatal cerebral amyloid angiopathy in early adult life. In all crystal structures of cystatin C studied to date, the protein has been found to form 3D domain-swapped dimers, created through a conformational change of a beta-hairpin loop, L1, from the papain-binding epitope. We have created monomer-stabilized human cystatin C, with an engineered disulfide bond (L47C)-(G69C) between the structural elements that become separated upon domain swapping. The mutant has drastically reduced dimerization and fibril formation properties, but its inhibition of papain is unaltered. The structure confirms the success of the protein engineering experiment to abolish 3D domain swapping and, in consequence, amyloid fibril formation. It illustrates for the first time the fold of monomeric cystatin C and allows verification of earlier predictions based on the domain-swapped forms and on the structure of chicken cystatin. Importantly, the structure defines the so-far unknown conformation of loop L1, which is essential for the inhibition of papain-like cysteine proteases.
Organizational Affiliation:
Faculty of Chemistry, Department of Crystallography, A. Mickiewicz University, Poznan, Poland.