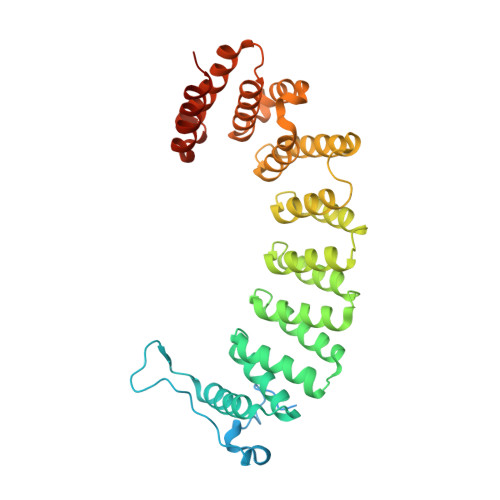
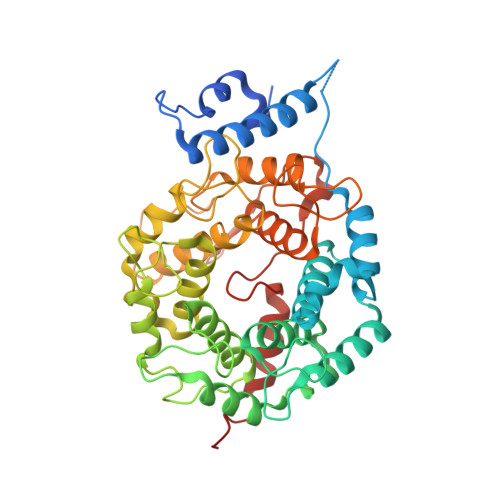
Analysis of the eukaryotic prenylome by isoprenoid affinity tagging
Nguyen, U.T.T., Guo, Z., Delon, C., Wu, Y., Deraeve, C., Franzel, B., Bon, R.S., Blankenfeldt, W., Goody, R.S., Waldmann, H., Wolters, D., Alexandrov, K.(2009) Nat Chem Biol 5: 227-235
- PubMed: 19219049
- DOI: https://doi.org/10.1038/nchembio.149
- Primary Citation of Related Structures:
3EU5, 3EUV - PubMed Abstract:
Protein prenylation is a widespread phenomenon in eukaryotic cells that affects many important signaling molecules. We describe the structure-guided design of engineered protein prenyltransferases and their universal synthetic substrate, biotin-geranylpyrophosphate. These new tools allowed us to detect femtomolar amounts of prenylatable proteins in cells and organs and to identify their cognate protein prenyltransferases. Using this approach, we analyzed the in vivo effects of protein prenyltransferase inhibitors. Whereas some of the inhibitors displayed the expected activities, others lacked in vivo activity or targeted a broader spectrum of prenyltransferases than previously believed. To quantitate the in vivo effect of the prenylation inhibitors, we profiled biotin-geranyl-tagged RabGTPases across the proteome by mass spectrometry. We also demonstrate that sites of active vesicular transport carry most of the RabGTPases. This approach enables a quantitative proteome-wide analysis of the regulation of protein prenylation and its modulation by therapeutic agents.
Organizational Affiliation:
Max Planck Institute of Molecular Physiology, Dortmund, Germany.