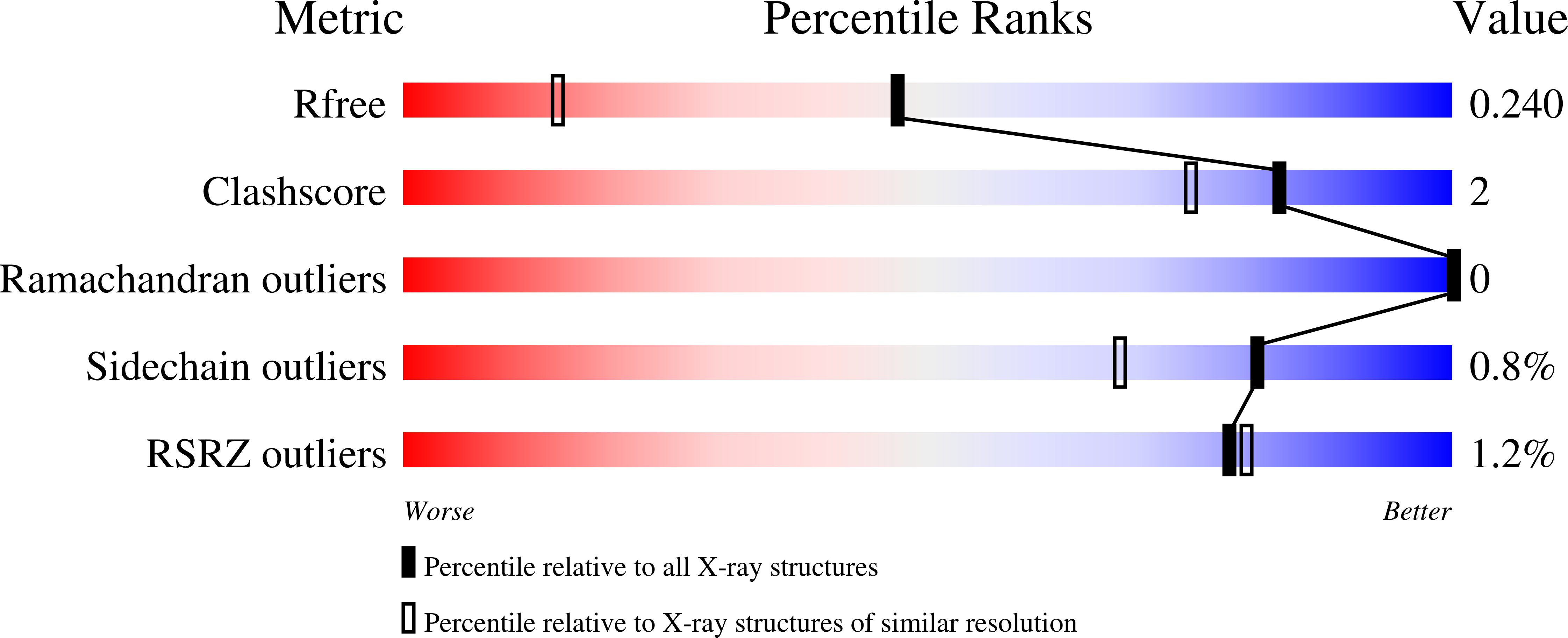
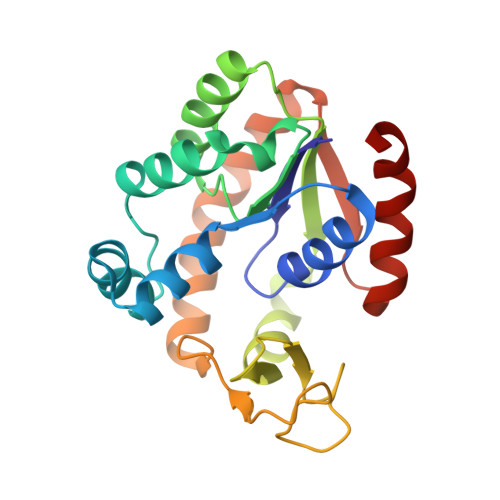
Effectiveness and limitations of local structural entropy optimization in the thermal stabilization of mesophilic and thermophilic adenylate kinases.
Moon, S., Bannen, R.M., Rutkoski, T.J., Phillips, G.N., Bae, E.(2014) Proteins 82: 2631-2642
- PubMed: 24931334
- DOI: https://doi.org/10.1002/prot.24627
- Primary Citation of Related Structures:
3DL0, 4QBF, 4QBG, 4QBH, 4QBI - PubMed Abstract:
Local structural entropy (LSE) is a descriptor for the extent of conformational heterogeneity in short protein sequences that is computed from structural information derived from the Protein Data Bank. Reducing the LSE of a protein sequence by introducing amino acid mutations can result in fewer conformational states and thus a more stable structure, indicating that LSE optimization can be used as a protein stabilization method. Here, we describe a series of LSE optimization experiments designed to stabilize mesophilic and thermophilic adenylate kinases (AKs) and report crystal structures of LSE-optimized AK variants. In the mesophilic AK, thermal stabilization by LSE reduction was effective but limited. Structural analyses of the LSE-optimized mesophilic AK variants revealed a strong correlation between LSE and the apolar buried surface area. Additional mutations designed to introduce noncovalent interactions between distant regions of the polypeptide resulted in further stabilization. Unexpectedly, optimizing the LSE of the thermophilic AK resulted in a decrease in thermal stability. This destabilization was reduced when charged residues were excluded from the possible substitutions during LSE optimization. These observations suggest that stabilization by LSE reduction may result from the optimization of local hydrophobic contacts. The limitations of this process are likely due to ignorance of other interactions that bridge distant regions in a given amino acid sequence. Our results illustrate the effectiveness and limitations of LSE optimization as a protein stabilization strategy and highlight the importance and complementarity of local conformational stability and global interactions in protein thermal stability.
Organizational Affiliation:
Department of Agricultural Biotechnology, Seoul National University, Seoul, 151-921, Korea.