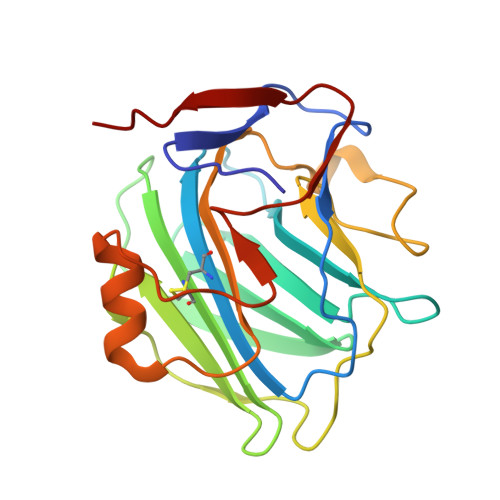
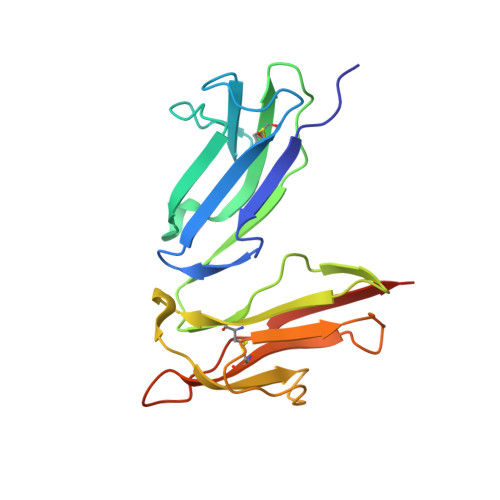
Structural recognition and functional activation of FcgammaR by innate pentraxins.
Lu, J., Marnell, L.L., Marjon, K.D., Mold, C., Du Clos, T.W., Sun, P.D.(2008) Nature 456: 989-992
- PubMed: 19011614
- DOI: https://doi.org/10.1038/nature07468
- Primary Citation of Related Structures:
3D5O - PubMed Abstract:
Pentraxins are a family of ancient innate immune mediators conserved throughout evolution. The classical pentraxins include serum amyloid P component (SAP) and C-reactive protein, which are two of the acute-phase proteins synthesized in response to infection. Both recognize microbial pathogens and activate the classical complement pathway through C1q (refs 3 and 4). More recently, members of the pentraxin family were found to interact with cell-surface Fcgamma receptors (FcgammaR) and activate leukocyte-mediated phagocytosis. Here we describe the structural mechanism for pentraxin's binding to FcgammaR and its functional activation of FcgammaR-mediated phagocytosis and cytokine secretion. The complex structure between human SAP and FcgammaRIIa reveals a diagonally bound receptor on each SAP pentamer with both D1 and D2 domains of the receptor contacting the ridge helices from two SAP subunits. The 1:1 stoichiometry between SAP and FcgammaRIIa infers the requirement for multivalent pathogen binding for receptor aggregation. Mutational and binding studies show that pentraxins are diverse in their binding specificity for FcgammaR isoforms but conserved in their recognition structure. The shared binding site for SAP and IgG results in competition for FcgammaR binding and the inhibition of immune-complex-mediated phagocytosis by soluble pentraxins. These results establish antibody-like functions for pentraxins in the FcgammaR pathway, suggest an evolutionary overlap between the innate and adaptive immune systems, and have new therapeutic implications for autoimmune diseases.
Organizational Affiliation:
Structural Immunology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Rockville, Maryland 20852, USA.