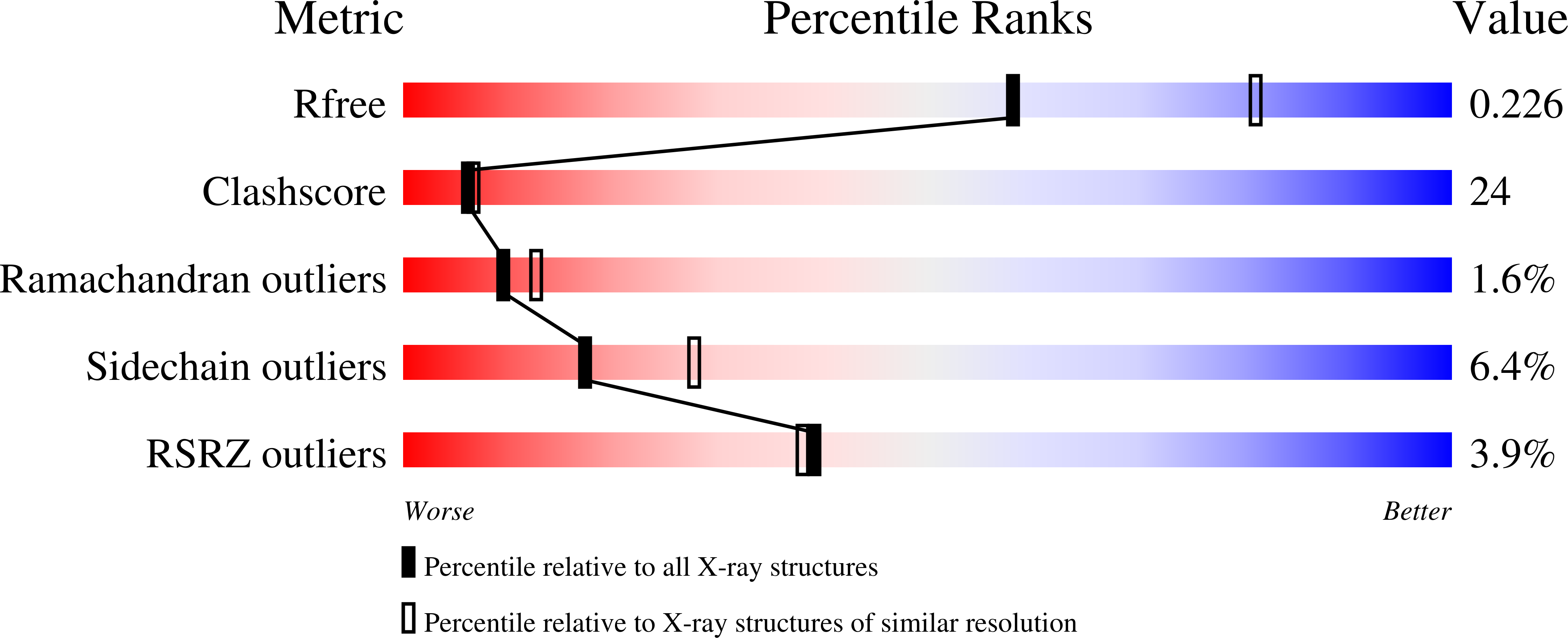
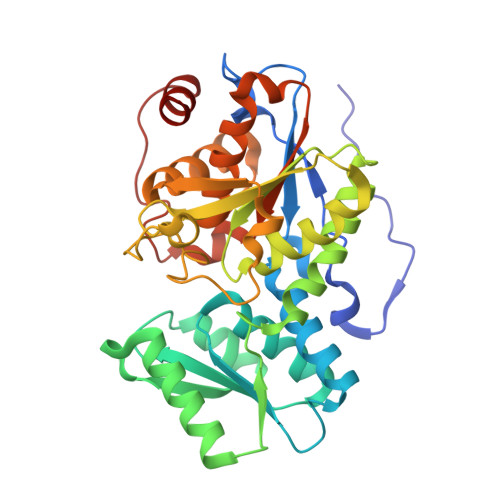
Crystal structure of native O-acetyl-serine sulfhydrylase from Entamoeba histolytica and its complex with cysteine: structural evidence for cysteine binding and lack of interactions with serine acetyl transferase.
Chinthalapudi, K., Kumar, M., Kumar, S., Jain, S., Alam, N., Gourinath, S.(2008) Proteins 72: 1222-1232
- PubMed: 18350570
- DOI: https://doi.org/10.1002/prot.22013
- Primary Citation of Related Structures:
2PQM, 3BM5 - PubMed Abstract:
Cysteine plays a major role in the antioxidative defense mechanisms of the human parasite Entameoba histolytica. The major route of cysteine biosynthesis in this parasite is the condensation of O-acetylserine with sulfide by the de novo cysteine biosynthetic pathway involving two key enzymes O-acetyl-L-serine sulfhydrylase (OASS) and serine acetyl transferase (SAT). The crystal structure of native OASS from Entameoba histolytica (EhOASS) has been determined at 1.86 A resolution and in complex with its product cysteine at 2.4 A resolution. In comparison with other known OASS structures, insertion in the N-terminal region and C-terminal helix reveal critical differences, which may influence the protein-protein interactions. In spite of lacking chloride binding site at the dimeric interface, the N-terminal extension compared with other known cysteine synthases, participates in dimeric interactions in an interesting domain swapping manner, enabling it to form a stronger dimer. Sulfate is bound in the active site of the native structure, which is replaced by cysteine in the cysteine bound form causing reorientation of the small N-terminal domain and thus closure of the active site. Ligand binding constants of OAS, Cys, and Met with EhOASS are comparable with other known OASS indicating similar active site arrangement and dynamics. The cysteine complexed structure represents the snapshot of the enzyme just before releasing the final product with a closed active site. The C-terminal helix positioning in the EhOASS may effect its interactions with EhSAT and thus influencing the formation of the cysteine synthase complex in this organism.
Organizational Affiliation:
School of Life Sciences, Jawaharlal Nehru University, New Delhi 110067, India.