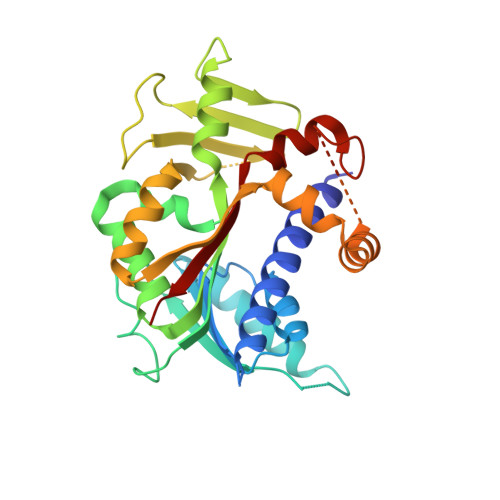
The crystal structure of human RNA (guanine-7-) methyltransferase in complex with SAH.
Wu, H., Lunin, V.V., Zeng, H., Antoshenko, T., MacKenzie, F., Weigelt, J., Arrowsmith, C.H., Edwards, A.M., Bochkarev, A., Min, J., Plotnikov, A.N.To be published.