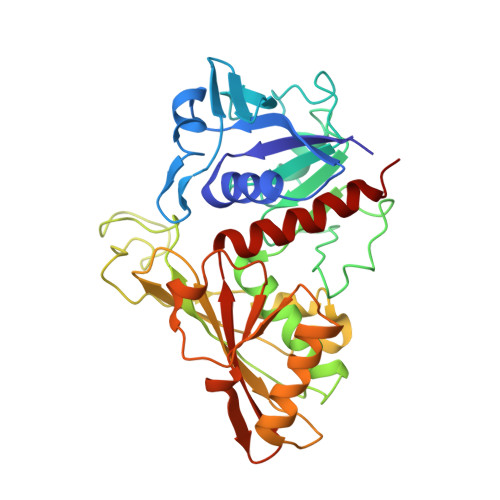
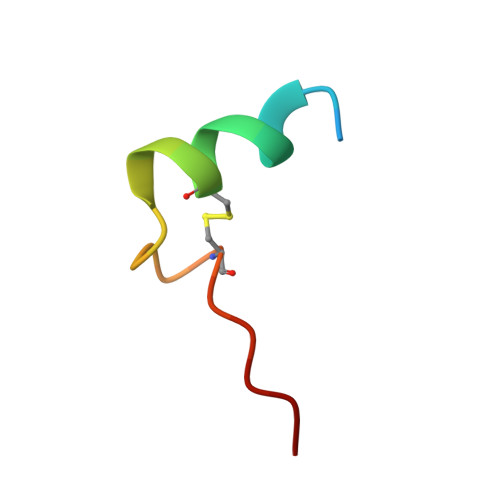
Structure Basis for the Regulation of Glyceraldehyde-3-Phosphate Dehydrogenase Activity via the Intrinsically Disordered Protein CP12.
Matsumura, H., Kai, A., Maeda, T., Tamoi, M., Satoh, A., Tamura, H., Hirose, M., Ogawa, T., Kizu, N., Wadano, A., Inoue, T., Shigeoka, S.(2011) Structure 19: 1846-1854
- PubMed: 22153507
- DOI: https://doi.org/10.1016/j.str.2011.08.016
- Primary Citation of Related Structures:
3B1J, 3B1K, 3B20 - PubMed Abstract:
The reversible formation of a glyceraldehyde-3-phosphate dehydrogenase (GAPDH)-CP12-phosphoribulokinase (PRK) supramolecular complex, identified in oxygenic photosynthetic organisms, provides light-dependent Calvin cycle regulation in a coordinated manner. An intrinsically disordered protein (IDP) CP12 acts as a linker to sequentially bind GAPDH and PRK to downregulate both enzymes. Here, we report the crystal structures of the ternary GAPDH-CP12-NAD and binary GAPDH-NAD complexes from Synechococcus elongates. The GAPDH-CP12 complex structure reveals that the oxidized CP12 becomes partially structured upon GAPDH binding. The C-terminus of CP12 is inserted into the active-site region of GAPDH, resulting in competitive inhibition of GAPDH. This study also provides insight into how the GAPDH-CP12 complex is dissociated by a high NADP(H)/NAD(H) ratio. An unexpected increase in negative charge potential that emerged upon CP12 binding highlights the biological function of CP12 in the sequential assembly of the supramolecular complex.
Organizational Affiliation:
Department of Applied Chemistry, Graduate School of Engineering, Osaka University, 2-1 Yamadaoka, Suita, Osaka 565-0871, Japan. matsumura@chem.eng.osaka-u.ac.jp