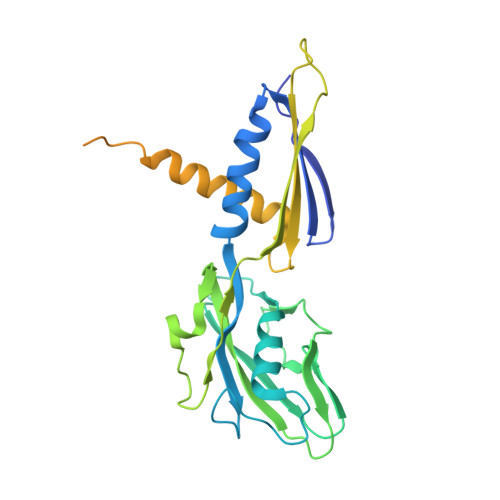
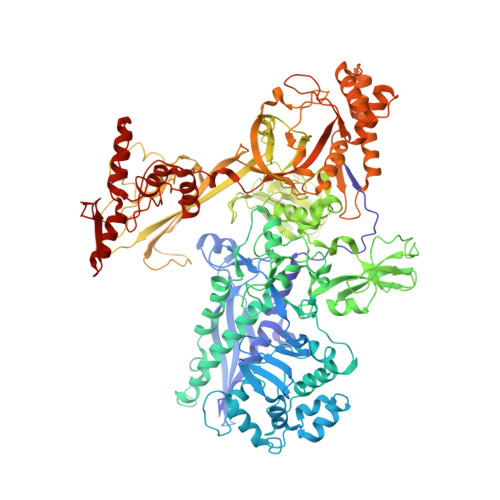
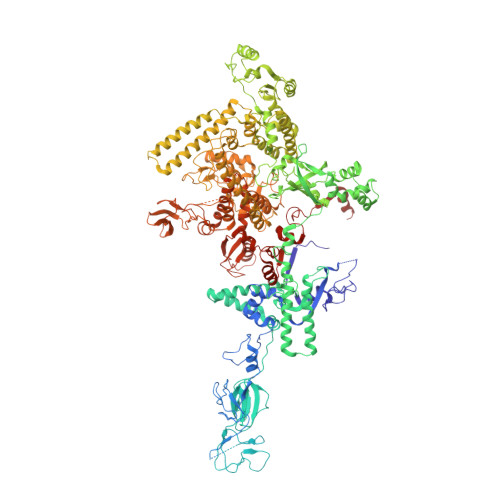
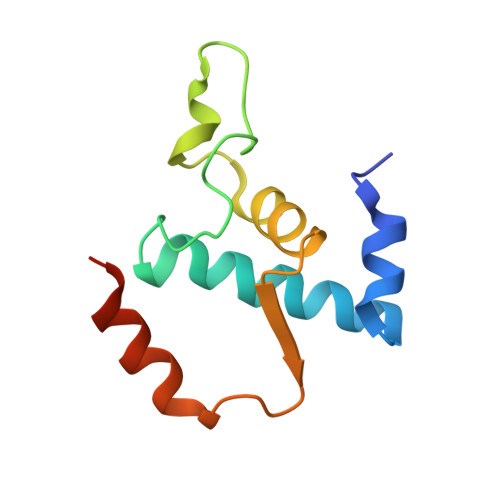
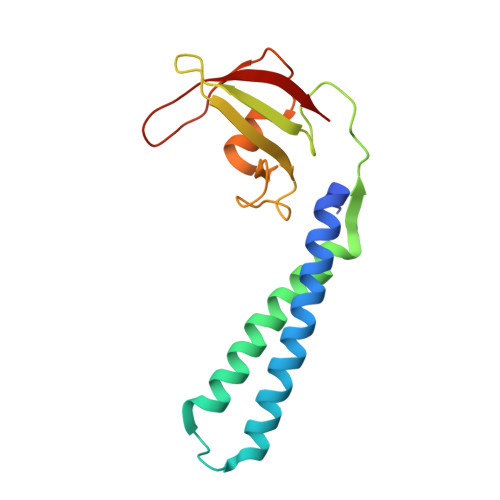
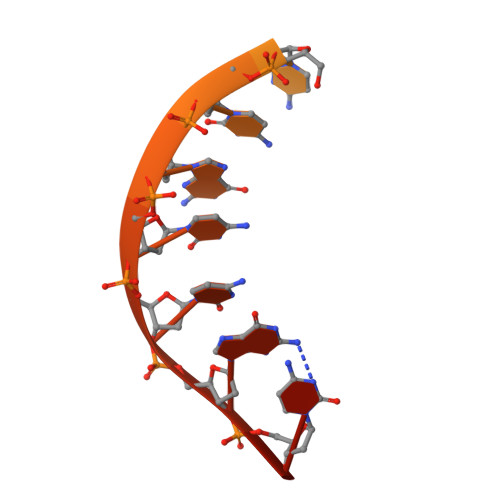
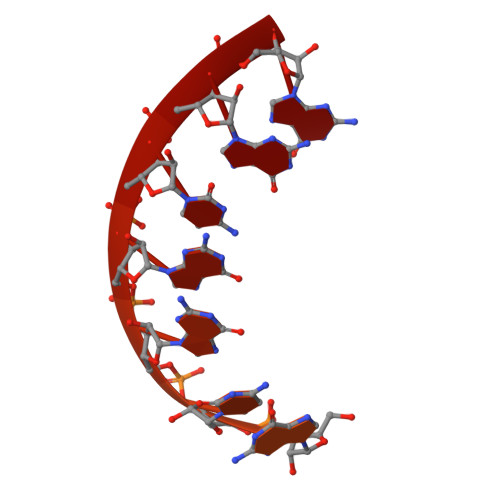
Crystal structure of bacterial RNA polymerase bound with a transcription inhibitor protein
Tagami, S., Sekine, S.I., Kumarevel, T., Hino, N., Murayama, Y., Kamegamori, S., Yamamoto, M., Sakamoto, K., Yokoyama, S.(2010) Nature 468: 978-982
- PubMed: 21124318
- DOI: https://doi.org/10.1038/nature09573
- Primary Citation of Related Structures:
3AOH, 3AOI - PubMed Abstract:
The multi-subunit DNA-dependent RNA polymerase (RNAP) is the principal enzyme of transcription for gene expression. Transcription is regulated by various transcription factors. Gre factor homologue 1 (Gfh1), found in the Thermus genus, is a close homologue of the well-conserved bacterial transcription factor GreA, and inhibits transcription initiation and elongation by binding directly to RNAP. The structural basis of transcription inhibition by Gfh1 has remained elusive, although the crystal structures of RNAP and Gfh1 have been determined separately. Here we report the crystal structure of Thermus thermophilus RNAP complexed with Gfh1. The amino-terminal coiled-coil domain of Gfh1 fully occludes the channel formed between the two central modules of RNAP; this channel would normally be used for nucleotide triphosphate (NTP) entry into the catalytic site. Furthermore, the tip of the coiled-coil domain occupies the NTP β-γ phosphate-binding site. The NTP-entry channel is expanded, because the central modules are 'ratcheted' relative to each other by ∼7°, as compared with the previously reported elongation complexes. This 'ratcheted state' is an alternative structural state, defined by a newly acquired contact between the central modules. Therefore, the shape of Gfh1 is appropriate to maintain RNAP in the ratcheted state. Simultaneously, the ratcheting expands the nucleic-acid-binding channel, and kinks the bridge helix, which connects the central modules. Taken together, the present results reveal that Gfh1 inhibits transcription by preventing NTP binding and freezing RNAP in the alternative structural state. The ratcheted state might also be associated with other aspects of transcription, such as RNAP translocation and transcription termination.
Organizational Affiliation:
Department of Biophysics and Biochemistry, Graduate School of Science, University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.