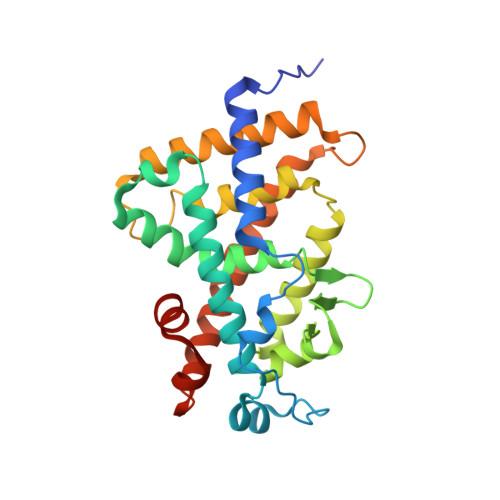
Structure-function relationships and crystal structures of the vitamin D receptor bound 2 alpha-methyl-(20S,23S)- and 2 alpha-methyl-(20S,23R)-epoxymethano-1 alpha,25-dihydroxyvitamin D3
Antony, P., Sigueiro, R., Huet, T., Sato, Y., Ramalanjaona, N., Rodrigues, L.C., Mourino, A., Moras, D., Rochel, N.(2010) J Med Chem 53: 1159-1171
- PubMed: 20070104
- DOI: https://doi.org/10.1021/jm9014636
- Primary Citation of Related Structures:
3A3Z, 3A40 - PubMed Abstract:
The vitamin D nuclear receptor is a ligand-dependent transcription factor that controls multiple biological responses such as cell proliferation, immune responses, and bone mineralization. Numerous 1 alpha,25(OH)(2)D(3) analogues, which exhibit low calcemic side effects and/or antitumoral properties, have been synthesized. We recently showed that the synthetic analogue (20S,23S)-epoxymethano-1 alpha,25-dihydroxyvitamin D(3) (2a) acts as a 1 alpha,25(OH)(2)D(3) superagonist and exhibits both antiproliferative and prodifferentiating properties in vitro. Using this information and on the basis of the crystal structures of human VDR ligand binding domain (hVDR LBD) bound to 1 alpha,25(OH)(2)D(3), 2 alpha-methyl-1 alpha,25(OH)(2)D(3), or 2a, we designed a novel analogue, 2 alpha-methyl-(20S,23S)-epoxymethano-1 alpha,25-dihydroxyvitamin D(3) (4a), in order to increase its transactivation potency. Here, we solved the crystal structures of the hVDR LBD in complex with the 4a (C23S) and its epimer 4b (C23R) and determined their correlation with specific biological outcomes.
Organizational Affiliation:
Departement de Biologie et de Génomique Structurales, Centre National de la Recherche Scientifique, Institut National de la Santé de la Recherche Médicale, Université de Strasbourg, 1 Rue Laurent Fries, 67404 Illkirch, France.