Novel Nadph-Cysteine Covalent Adduct Found in the Active Site of an Aldehyde Dehydrogenase.
Diaz-Sanchez, A.G., Gonzalez-Segura, L., Rudino-Pinera, E., Lira-Rocha, A., Torres-Larios, A., Munoz-Clares, R.A.(2011) Biochem J 439: 443
- PubMed: 21732915
- DOI: https://doi.org/10.1042/BJ20110376
- Primary Citation of Related Structures:
2WOX, 3ZQA - PubMed Abstract:
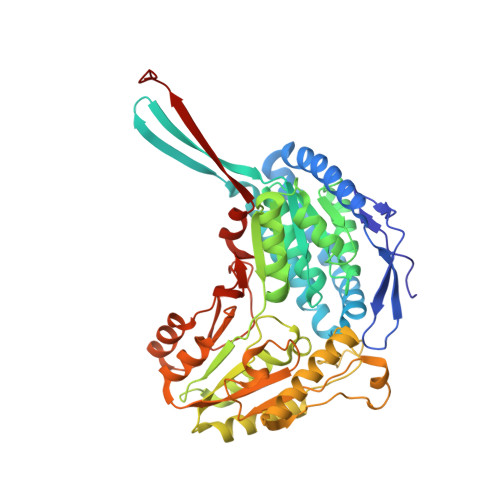
PaBADH (Pseudomonas aeruginosa betaine aldehyde dehydrogenase) catalyses the irreversible NAD(P)+-dependent oxidation of betaine aldehyde to its corresponding acid, the osmoprotector glycine betaine. This reaction is involved in the catabolism of choline and in the response of this important pathogen to the osmotic and oxidative stresses prevalent in infection sites. The crystal structure of PaBADH in complex with NADPH showed a novel covalent adduct between the C2N of the pyridine ring and the sulfur atom of the catalytic cysteine residue, Cys286. This kind of adduct has not been reported previously either for a cysteine residue or for a low-molecular-mass thiol. The Michael addition of the cysteine thiolate in the 'resting' conformation to the double bond of the α,β-unsaturated nicotinamide is facilitated by the particular conformation of NADPH in the active site of PaBADH (also observed in the crystal structure of the Cys286Ala mutant) and by an ordered water molecule hydrogen bonded to the carboxamide group. Reversible formation of NAD(P)H-Cys286 adducts in solution causes reversible enzyme inactivation as well as the loss of Cys286 reactivity towards thiol-specific reagents. This novel covalent modification may provide a physiologically relevant regulatory mechanism of the irreversible PaBADH-catalysed reaction, preventing deleterious decreases in the intracellular NAD(P)+/NAD(P)H ratios.
Organizational Affiliation:
Departamento de Bioquímica, Universidad Nacional Autónoma de México, Ciudad Universitaria. clares@servidor.unam.mx