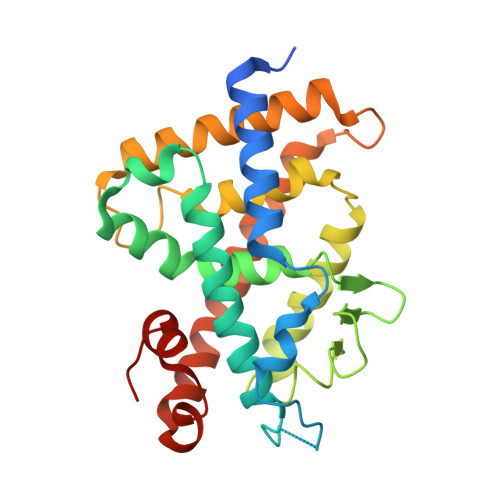
Crystal structures of complexes of vitamin D receptor ligand-binding domain with lithocholic acid derivatives.
Masuno, H., Ikura, T., Morizono, D., Orita, I., Yamada, S., Shimizu, M., Ito, N.(2013) J Lipid Res 54: 2206-2213
- PubMed: 23723390
- DOI: https://doi.org/10.1194/jlr.M038307
- Primary Citation of Related Structures:
3W5P, 3W5Q, 3W5R, 3W5T - PubMed Abstract:
The secondary bile acid lithocholic acid (LCA) and its derivatives act as selective modulators of the vitamin D receptor (VDR), although their structures fundamentally differ from that of the natural hormone 1α,25-dihydroxyvitamin D3 [1,25(OH)2D3)]. Here, we have determined the crystal structures of the ligand-binding domain of rat VDR (VDR-LBD) in ternary complexes with a synthetic partial peptide of the coactivator MED1 (mediator of RNA polymerase II transcription subunit 1) and four ligands, LCA, 3-keto LCA, LCA acetate, and LCA propionate, with the goal of elucidating their agonistic mechanism. LCA and its derivatives bind to the same ligand-binding pocket (LBP) of VDR-LBD that 1,25(OH)2D3 binds to, but in the opposite orientation; their A-ring is positioned at the top of the LBP, whereas their acyclic tail is located at the bottom of the LBP. However, most of the hydrophobic and hydrophilic interactions observed in the complex with 1,25(OH)2D3 are reproduced in the complexes with LCA and its derivatives. Additional interactions between VDR-LBD and the C-3 substituents of the A-ring are also observed in the complexes with LCA and its derivatives. These may result in the observed difference in the potency among the LCA-type ligands.
Organizational Affiliation:
Institute of Biomaterials and Bioengineering, Tokyo Medical and Dental University, Chiyoda-ku, Tokyo 101-0062, Japan.