Design and synthesis of a series of alpha-benzyl phenylpropanoic acid-type peroxisome proliferator-activated receptor (PPAR) gamma partial agonists with improved aqueous solubility
Ohashi, M., Oyama, T., Putranto, E.W., Waku, T., Nobusada, H., Kataoka, K., Matsuno, K., Yashiro, M., Morikawa, K., Huh, N.H., Miyachi, H.(2013) Bioorg Med Chem 21: 2319-2332
- PubMed: 23490155
- DOI: https://doi.org/10.1016/j.bmc.2013.02.003
- Primary Citation of Related Structures:
3VSO - PubMed Abstract:
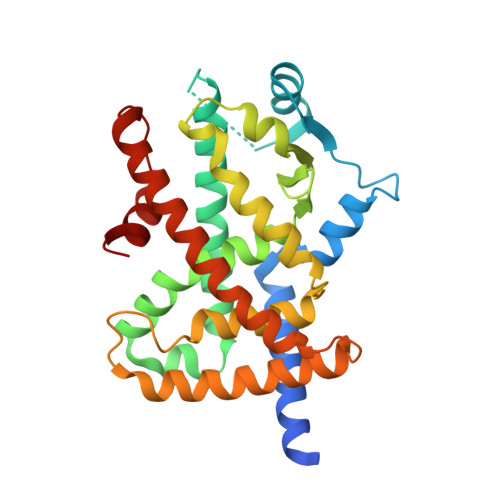
In the continuing study directed toward the development of peroxisome proliferator-activated receptor gamma (hPPARγ) agonist, we attempted to improve the water solubility of our previously developed hPPARγ-selective agonist 3, which is insufficiently soluble for practical use, by employing two strategies: introducing substituents to reduce its molecular planarity and decreasing its hydrophobicity via replacement of the adamantyl group with a heteroaromatic ring. The first approach proved ineffective, but the second was productive. Here, we report the design and synthesis of a series of α-benzyl phenylpropanoic acid-type hPPARγ partial agonists with improved aqueous solubility. Among them, we selected (R)-7j, which activates hPPARγ to the extent of about 65% of the maximum observed with a full agonist, for further evaluation. The ligand-binding mode and the reason for the partial-agonistic activity are discussed based on X-ray-determined structure of the complex of hPPARγ ligand-binding domain (LBD) and (R)-7j with previously reported ligand-LDB structures. Preliminal apoptotic effect of (R)-7j against human scirrhous gastric cancer cell line OCUM-2MD3 is also described.
Organizational Affiliation:
Graduate School of Medicine, Dentistry and Pharmaceutical Sciences, Okayama University, 1-1-1 Tsushima-Naka, Kita-ku, Okayama 700-8530, Japan.