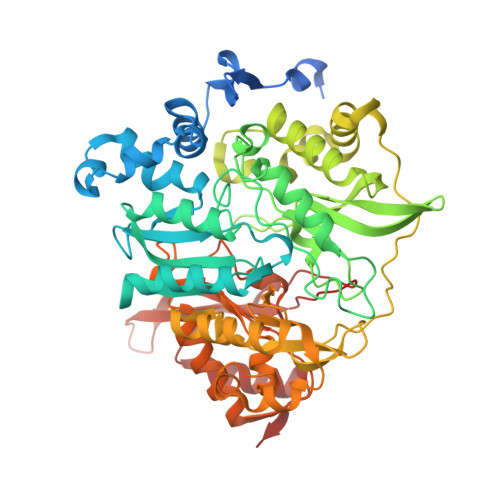
X-ray crystal structure of a mutant assimilatory nitrite reductase that shows sulfite reductase-like activity
Nakano, S., Takahashi, M., Sakamoto, A., Morikawa, H., Katayanagi, K.(2012) Chem Biodivers 9: 1989-1999
- PubMed: 22976986
- DOI: https://doi.org/10.1002/cbdv.201100442
- Primary Citation of Related Structures:
3VLX, 3VLY, 3VLZ, 3VM0, 3VM1 - PubMed Abstract:
Assimilatory nitrite reductase (aNiR) reduces nitrite ions (NO(2)(-)) to ammonium ions (NH(4)(+)), whereas assimilatory sulfite reductase reduces sulfite (SO(3)(2-)) to hydrogen sulfide (HS(-)). Although aNiR can also reduce SO(3)(2-), its activity is much lower than when NO(2)(-) is reduced as the substrate. To increase the SO(3)(2-)-reduction activity of aNiR, we performed a N226K mutation of Nii3, a representative aNiR. The resulting Nii3-N226K variant could bind non-native targets, SO(3)(2-), and HCO(3)(-), in addition to its native target, i.e., NO(2)(-). We have determined the high-resolution structure of Nii3-N226K in its apo-state and in complex with SO(3)(2-), NO(2)(-), and HCO(3)(-). This analysis revealed conformational changes of Lys226 and the adjacent Lys224 upon binding of SO(3)(2-), but not NO(2)(-)In contrast, HCO(3)(-) binding induced a conformational change at Arg179. After replacing Asn226 with a positively charged Lys, aNiR showed affinity for several anions. A comparison of all ligand-bound structures for Nii3-N226K revealed that structural changes in the active site depend on the size of the substrate.
Organizational Affiliation:
Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, 1-3-1, Kagamiyama, Higashi-Hiroshima 739-8526, Japan.