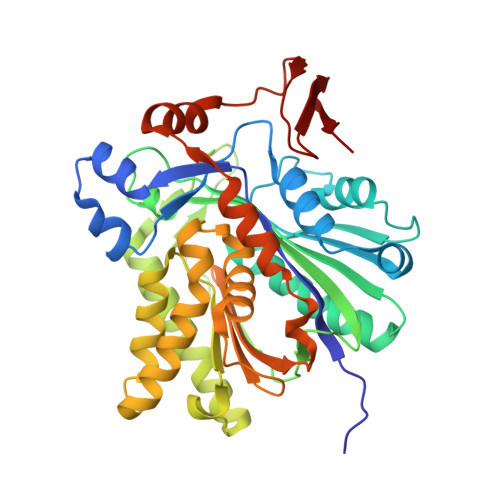
Biochemical and structural basis for inhibition of Enterococcus faecalis hydroxymethylglutaryl-CoA synthase, mvaS, by hymeglusin.
Skaff, D.A., Ramyar, K.X., McWhorter, W.J., Barta, M.L., Geisbrecht, B.V., Miziorko, H.M.(2012) Biochemistry 51: 4713-4722
- PubMed: 22510038
- DOI: https://doi.org/10.1021/bi300037k
- Primary Citation of Related Structures:
3V4N, 3V4X - PubMed Abstract:
Hymeglusin (1233A, F244, L-659-699) is established as a specific β-lactone inhibitor of eukaryotic hydroxymethylglutaryl-CoA synthase (HMGCS). Inhibition results from formation of a thioester adduct to the active site cysteine. In contrast, the effects of hymeglusin on bacterial HMG-CoA synthase, mvaS, have been minimally characterized. Hymeglusin blocks growth of Enterococcus faecalis. After removal of the inhibitor from culture media, a growth curve inflection point at 3.1 h is observed (vs 0.7 h for the uninhibited control). Upon hymeglusin inactivation of purified E. faecalis mvaS, the thioester adduct is more stable than that measured for human HMGCS. Hydroxylamine cleaves the thioester adduct; substantial enzyme activity is restored at a rate that is 8-fold faster for human HMGCS than for mvaS. Structural results explain these differences in enzyme-inhibitor thioester adduct stability and solvent accessibility. The E. faecalis mvaS-hymeglusin cocrystal structure (1.95 Å) reveals virtually complete occlusion of the bound inhibitor in a narrow tunnel that is largely sequestered from bulk solvent. In contrast, eukaryotic (Brassica juncea) HMGCS binds hymeglusin in a more solvent-exposed cavity.
Organizational Affiliation:
University of Missouri, Kansas City, MO, USA.