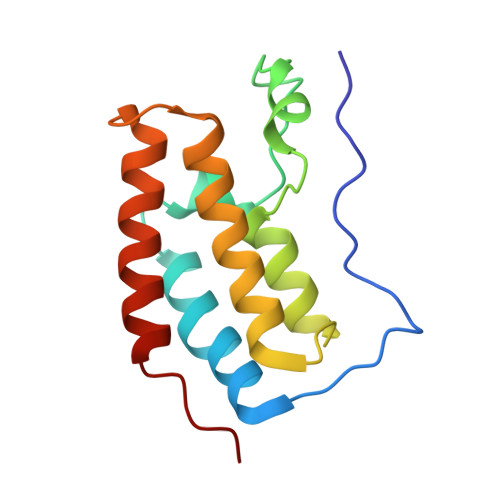
Benzodiazepines and benzotriazepines as protein interaction inhibitors targeting bromodomains of the BET family.
Filippakopoulos, P., Picaud, S., Fedorov, O., Keller, M., Wrobel, M., Morgenstern, O., Bracher, F., Knapp, S.(2012) Bioorg Med Chem 20: 1878-1886
- PubMed: 22137933
- DOI: https://doi.org/10.1016/j.bmc.2011.10.080
- Primary Citation of Related Structures:
3U5J, 3U5K, 3U5L - PubMed Abstract:
Benzodiazepines are psychoactive drugs with anxiolytic, sedative, skeletal muscle relaxant and amnestic properties. Recently triazolo-benzodiazepines have been also described as potent and highly selective protein interaction inhibitors of bromodomain and extra-terminal (BET) proteins, a family of transcriptional co-regulators that play a key role in cancer cell survival and proliferation, but the requirements for high affinity interaction of this compound class with bromodomains has not been described. Here we provide insight into the structure-activity relationship (SAR) and selectivity of this versatile scaffold. In addition, using high resolution crystal structures we compared the binding mode of a series of benzodiazepine (BzD) and related triazolo-benzotriazepines (BzT) derivatives including clinically approved drugs such as alprazolam and midazolam. Our analysis revealed the importance of the 1-methyl triazolo ring system for BET binding and suggests modifications for the development of further high affinity bromodomain inhibitors.
Organizational Affiliation:
University of Oxford, Nuffield Department of Clinical Medicine, Structural Genomics Consortium, Old Road Campus Research Building, Roosevelt Drive, Oxford OX3 7LD, UK.