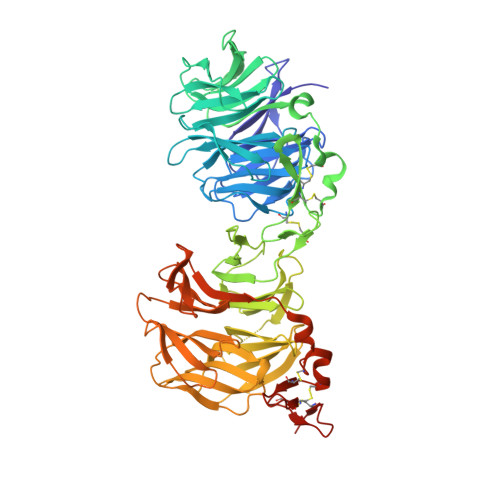
Structural Basis of Wnt Signaling Inhibition by Dickkopf Binding to LRP5/6.
Ahn, V.E., Chu, M.L., Choi, H.J., Tran, D., Abo, A., Weis, W.I.(2011) Dev Cell 21: 862-873
- PubMed: 22000856
- DOI: https://doi.org/10.1016/j.devcel.2011.09.003
- Primary Citation of Related Structures:
3S2K - PubMed Abstract:
LDL receptor-related proteins 5 and 6 (LRP5/6) are coreceptors for Wnt growth factors, and also bind Dkk proteins, secreted inhibitors of Wnt signaling. The LRP5/6 ectodomain contains four β-propeller/EGF-like domain repeats. The first two repeats, LRP6(1-2), bind to several Wnt variants, whereas LRP6(3-4) binds other Wnts. We present the crystal structure of the Dkk1 C-terminal domain bound to LRP6(3-4), and show that the Dkk1 N-terminal domain binds to LRP6(1-2), demonstrating that a single Dkk1 molecule can bind to both portions of the LRP6 ectodomain and thereby inhibit different Wnts. Small-angle X-ray scattering analysis of LRP6(1-4) bound to a noninhibitory antibody fragment or to full-length Dkk1 shows that in both cases the ectodomain adopts a curved conformation that places the first three repeats at a similar height relative to the membrane. Thus, Wnts bound to either portion of the LRP6 ectodomain likely bear a similar spatial relationship to Frizzled coreceptors.
Organizational Affiliation:
Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305, USA.