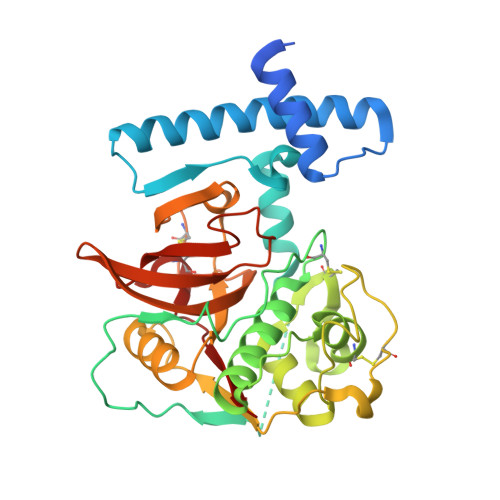
The 3D structure and function of digestive cathepsin L-like proteinases of Tenebrio molitor larval midgut.
Beton, D., Guzzo, C.R., Ribeiro, A.F., Farah, C.S., Terra, W.R.(2012) Insect Biochem Mol Biol 42: 655-664
- PubMed: 22659439
- DOI: https://doi.org/10.1016/j.ibmb.2012.04.010
- Primary Citation of Related Structures:
3QJ3, 3QT4 - PubMed Abstract:
Cathepsin L-like proteinases (CAL) are major digestive proteinases in the beetle Tenebrio molitor. Procathepsin Ls 2 (pCAL2) and 3 (pCAL3) were expressed as recombinant proteins in Escherichia coli, purified and activated under acidic conditions. Immunoblot analyses of different T. molitor larval tissues demonstrated that a polyclonal antibody to pCAL3 recognized pCAL3 and cathepsin L 3 (CAL3) only in the anterior two-thirds of midgut tissue and midgut luminal contents of T. molitor larvae. Furthermore, immunocytolocalization data indicated that pCAL3 occurs in secretory vesicles and microvilli in anterior midgut. Therefore CAL3, like cathepsin L 2 (CAL2), is a digestive enzyme secreted by T. molitor anterior midgut. CAL3 hydrolyses Z-FR-MCA and Z-RR-MCA (typical cathepsin substrates), whereas CAL2 hydrolyses only Z-FR-MCA. Active site mutants (pCAL2C25S and pCAL3C26S) were constructed by replacing the catalytic cysteine with serine to prevent autocatalytic processing. Recombinant pCAL2 and pCAL3 mutants (pCAL2C25S and pCAL3C26S) were prepared, crystallized and their 3D structures determined at 1.85 and 2.1 Å, respectively. While the overall structure of these enzymes is similar to other members of the papain superfamily, structural differences in the S2 subsite explain their substrate specificities. The data also supported models for CAL trafficking to lysosomes and to secretory vesicles to be discharged into midgut contents.
Organizational Affiliation:
Departamento de Bioquímica, Instituto de Química, Universidade de São Paulo, C. P. 26077, 05513-970 São Paulo, Brazil.