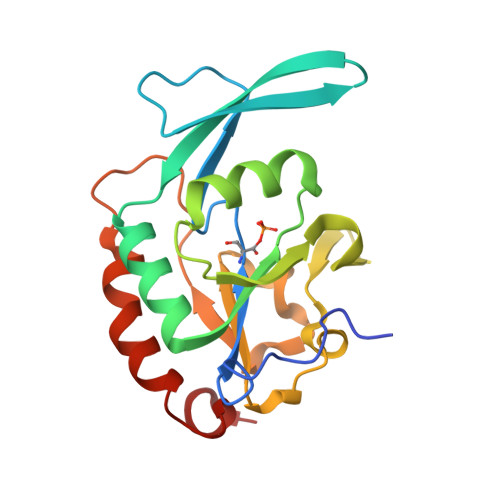
Structural and functional analysis of the phosphoryl transfer reaction mediated by the human small C-terminal domain phosphatase, Scp1.
Zhang, M., Liu, J., Kim, Y., Dixon, J.E., Pfaff, S.L., Gill, G.N., Noel, J.P., Zhang, Y.(2010) Protein Sci 19: 974-986
- PubMed: 20222012
- DOI: https://doi.org/10.1002/pro.375
- Primary Citation of Related Structures:
3L0B, 3L0C, 3L0Y - PubMed Abstract:
Human small C-terminal domain phosphatase 1 (Scp1) modulates the phosphorylation state of the C-terminal domain (CTD) of eukaryotic RNA polymerase II (RNAP II), with preference for phosphorylated Ser5 in the tandem heptad repeats of the CTD. Additionally, Scp1 was identified as a conserved regulator of neuronal stem cell development. Scp1 is a member of haloacid dehalogenase (HAD) superfamily, whose catalysis depends on a Mg(2+) ion and a DXDX(T/V) motif. The first Asp of the motif is identified as the nucleophile that is subject to phosphorylation leading to a phosphoryl-aspartate intermediate. This high-energy mixed anhydride intermediate is subsequently hydrolyzed to regenerate the enzyme. In the present study, we successfully captured the phosphoryl-aspartate intermediate in the crystal structure of a Scp1D206A mutant soaked with para-nitrophenyl phosphate (pNPP), providing strong evidence for the proposed mechanism. Furthermore, steady-state kinetic analysis of a variety of Scp1 mutants revealed the importance of Asp206 in Mg(2+) coordination mediated by a water molecule. Overall, we captured the snapshots of the phosphoryl transfer reaction at each stage of Scp1-mediated catalysis. Through structural-based sequence alignment, we show that the spatial position of the D206 side chain is strictly conserved throughout HAD family. Our results strongly suggest that Asp206 and its equivalent residues in other HAD family members play important structural and possible mechanistic roles.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Texas at Austin, Austin, Texas, USA.