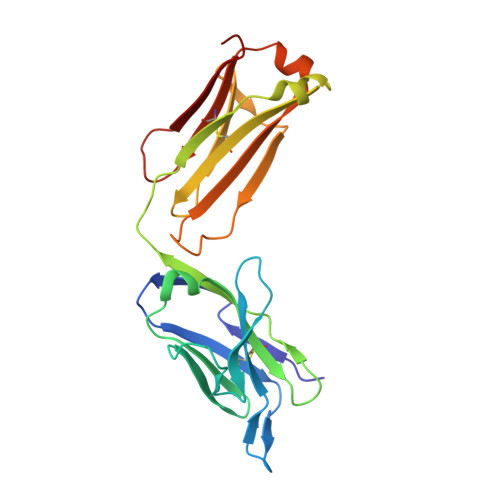
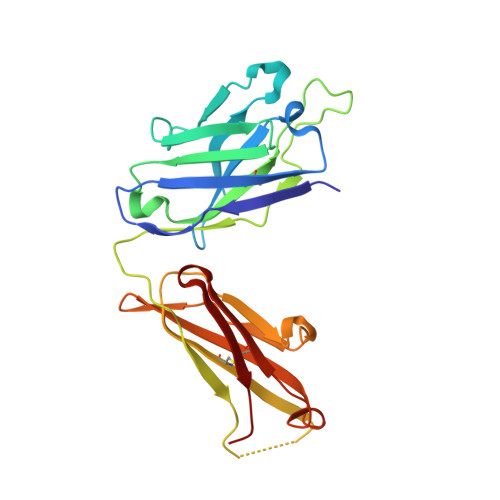
Antibodies raised against chlamydial lipopolysaccharide antigens reveal convergence in germline gene usage and differential epitope recognition
Brooks, C.L., Muller-Loennies, S., Borisova, S.N., Brade, L., Kosma, P., Hirama, T., Mackenzie, C.R., Brade, H., Evans, S.V.(2010) Biochemistry 49: 570-581
- PubMed: 20000757
- DOI: https://doi.org/10.1021/bi9011308
- Primary Citation of Related Structures:
3HZK, 3HZM, 3HZV, 3HZY, 3I02 - PubMed Abstract:
The structures of antigen-binding fragments from two related monoclonal antibodies have been determined to high resolution in the presence of several carbohydrate antigens raised against chlamydial lipopolysaccharide. With the exception of CDR H3, antibodies S54-10 and S73-2 are both derived from the same set of germline gene segments as the previously reported structures S25-2 and S45-18. Despite this similarity, the antibodies differ in specificity and the mechanism by which they recognize their cognate antigen. S54-10 uses an unrelated CDR H3 to recognize its antigen in a fashion analogous to S45-18; however, S73-2 recognizes the same antigen as S45-18 and S54-10 in a wholly unrelated manner. Together, these antibody-antigen structures provide snapshots into how the immune system uses the same set of inherited germline gene segments to generate multiple possible specificities that allow for differential recognition of epitopes and how unrelated CDR H3 sequences can result in convergent binding of clinically relevant bacterial antigens.
Organizational Affiliation:
University of Victoria, Department of Biochemistry and Microbiology, PO Box 3055 STN CSC, Victoria, BC, Canada V8P 3P6.