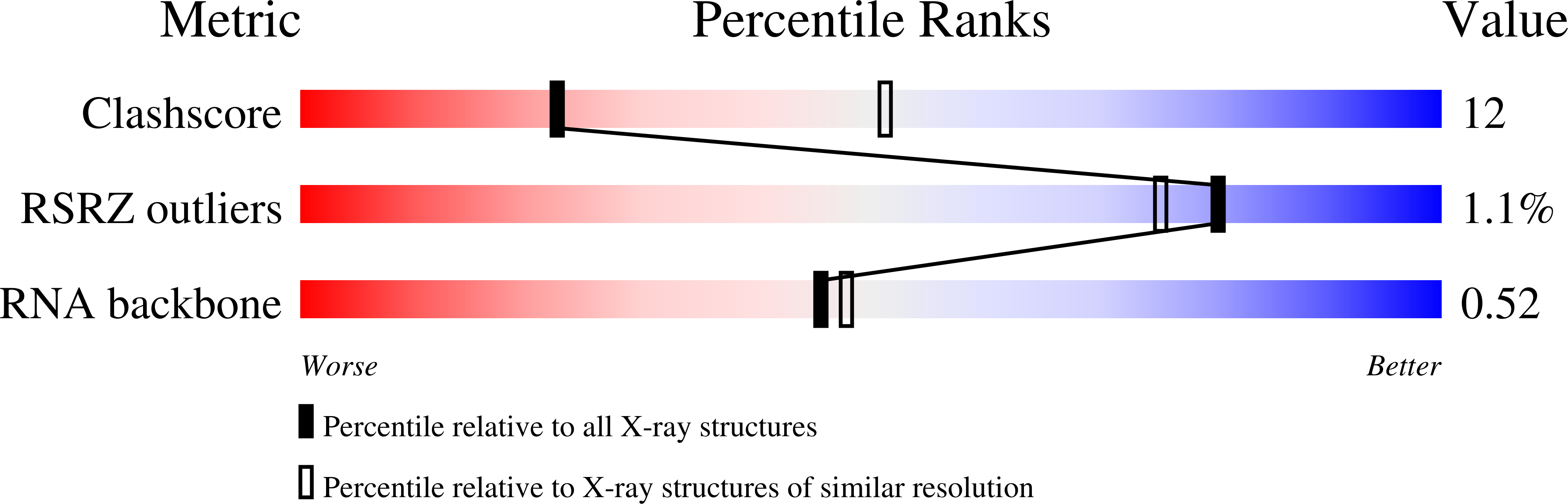Discrimination between Closely Related Cellular Metabolites by the SAM-I Riboswitch.
Montange, R.K., Mondragon, E., van Tyne, D., Garst, A.D., Ceres, P., Batey, R.T.(2010) J Mol Biol 396: 761-772
- PubMed: 20006621
- DOI: https://doi.org/10.1016/j.jmb.2009.12.007
- Primary Citation of Related Structures:
3GX2, 3GX3, 3GX5, 3GX6, 3GX7 - PubMed Abstract:
The SAM-I riboswitch is a cis-acting element of genetic control found in bacterial mRNAs that specifically binds S-adenosylmethionine (SAM). We previously determined the 2.9-A X-ray crystal structure of the effector-binding domain of this RNA element, revealing details of RNA-ligand recognition. To improve this structure, variations were made to the RNA sequence to alter lattice contacts, resulting in a 0.5-A improvement in crystallographic resolution and allowing for a more accurate refinement of the crystallographic model. The basis for SAM specificity was addressed by a structural analysis of the RNA complexed to S-adenosylhomocysteine (SAH) and sinefungin and by measuring the affinity of SAM and SAH for a series of mutants using isothermal titration calorimetry. These data illustrate the importance of two universally conserved base pairs in the RNA that form electrostatic interactions with the positively charged sulfonium group of SAM, thereby providing a basis for discrimination between SAM and SAH.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Colorado, Boulder, Campus Box 215, Boulder, CO 80309-0215, USA.