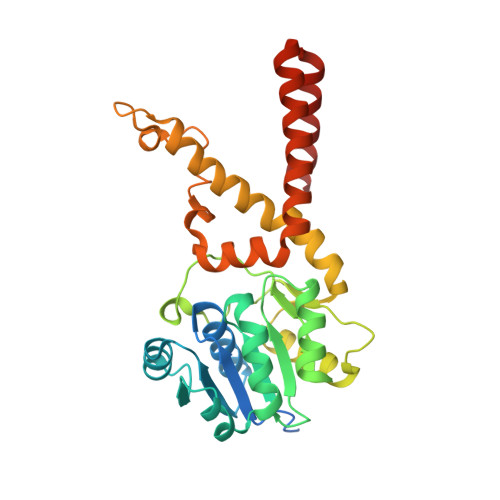
The Crystal Structure of Aquifex aeolicus Prephenate Dehydrogenase Reveals the Mode of Tyrosine Inhibition.
Sun, W., Shahinas, D., Bonvin, J., Hou, W., Kimber, M.S., Turnbull, J., Christendat, D.(2009) J Biol Chem 284: 13223-13232
- PubMed: 19279014
- DOI: https://doi.org/10.1074/jbc.M806272200
- Primary Citation of Related Structures:
3GGG, 3GGO, 3GGP - PubMed Abstract:
TyrA proteins belong to a family of dehydrogenases that are dedicated to l-tyrosine biosynthesis. The three TyrA subclasses are distinguished by their substrate specificities, namely the prephenate dehydrogenases, the arogenate dehydrogenases, and the cyclohexadienyl dehydrogenases, which utilize prephenate, l-arogenate, or both substrates, respectively. The molecular mechanism responsible for TyrA substrate selectivity and regulation is unknown. To further our understanding of TyrA-catalyzed reactions, we have determined the crystal structures of Aquifex aeolicus prephenate dehydrogenase bound with NAD(+) plus either 4-hydroxyphenylpyuvate, 4-hydroxyphenylpropionate, or l-tyrosine and have used these structures as guides to target active site residues for site-directed mutagenesis. From a combination of mutational and structural analyses, we have demonstrated that His-147 and Arg-250 are key catalytic and binding groups, respectively, and Ser-126 participates in both catalysis and substrate binding through the ligand 4-hydroxyl group. The crystal structure revealed that tyrosine, a known inhibitor, binds directly to the active site of the enzyme and not to an allosteric site. The most interesting finding though, is that mutating His-217 relieved the inhibitory effect of tyrosine on A. aeolicus prephenate dehydrogenase. The identification of a tyrosine-insensitive mutant provides a novel avenue for designing an unregulated enzyme for application in metabolic engineering.
Organizational Affiliation:
Department of Cell and Systems Biology, University of Toronto, Ontario, Canada.