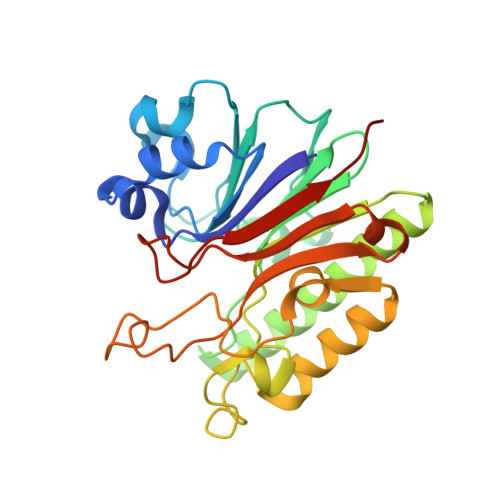
Crystal Structure Analysis of DNA Uridine Endonuclease Mth212 Bound to DNA
Lakomek, K., Dickmanns, A., Ciirdaeva, E., Schomacher, L., Ficner, R.(2010) J Mol Biol 399: 604-617
- PubMed: 20434457
- DOI: https://doi.org/10.1016/j.jmb.2010.04.044
- Primary Citation of Related Structures:
3FZI, 3G00, 3G0A, 3G0R, 3G1K, 3G2C, 3G2D, 3G38, 3G3C, 3G3Y, 3G4T, 3G8V, 3G91, 3GA6 - PubMed Abstract:
The reliable repair of pre-mutagenic U/G mismatches that originated from hydrolytic cytosine deamination is crucial for the maintenance of the correct genomic information. In most organisms, any uracil base in DNA is attacked by uracil DNA glycosylases (UDGs), but at least in Methanothermobacter thermautotrophicus DeltaH, an alternative strategy has evolved. The exonuclease III homologue Mth212 from the thermophilic archaeon M. thermautotrophicus DeltaH exhibits a DNA uridine endonuclease activity in addition to the apyrimidinic/apurinic site endonuclease and 3'-->5'exonuclease functions. Mth212 alone compensates for the lack of a UDG in a single-step reaction thus substituting the two-step pathway that requires the consecutive action of UDG and apyrimidinic/apurinic site endonuclease. In order to gain deeper insight into the structural basis required for the specific uridine recognition by Mth212, we have characterized the enzyme by means of X-ray crystallography. Structures of Mth212 wild-type or mutant proteins either alone or in complex with DNA substrates and products have been determined to a resolution of up to 1.2 A, suggesting key residues for the uridine endonuclease activity. The insertion of the side chain of Arg209 into the DNA helical base stack resembles interactions observed in human UDG and seems to be crucial for the uridine recognition. In addition, Ser171, Asn153, and Lys125 in the substrate binding pocket appear to have important functions in the discrimination of aberrant uridine against naturally occurring thymidine and cytosine residues in double-stranded DNA.
Organizational Affiliation:
Department of Molecular Structural Biology, Institute of Microbiology and Genetics, Georg-August University Göttingen, Justus-von-Liebig Weg 11, D-37077 Göttingen, Germany.