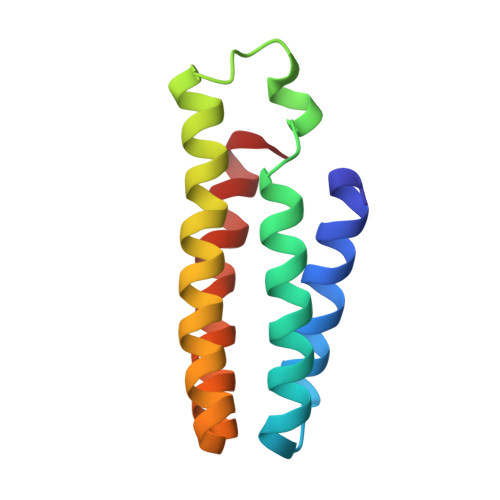
A superprotein triangle driven by nickel(II) coordination: exploiting non-natural metal ligands in protein self-assembly
Radford, R.J., Tezcan, F.A.(2009) J Am Chem Soc 131: 9136-9137
- PubMed: 19527025
- DOI: https://doi.org/10.1021/ja9000695
- Primary Citation of Related Structures:
3FOO, 3FOP - PubMed Abstract:
We previously devised a strategy (metal-directed protein self-assembly, MDPSA) that utilizes the simultaneous stability, lability, and directionality of metal-ligand bonds to drive protein-protein interactions. Here we show that both the structural and functional scopes of MDPSA can be broadened by incorporation of non-natural metal-chelating ligands onto protein surfaces. A cytochrome cb(562) variant, MBP-Phen1, which features a covalently attached phenanthroline (Phen) group on its surface, self-assembles into an unusual triangular architecture (Ni(3):MBP-Phen1(3)) upon binding Ni as a result of specific Phen-protein interactions. The crystal structure of Ni(3):MBP-Phen1(3) reveals that the Phen group is buried in a small pocket on the protein surface, which results in an unsaturated Ni coordination environment.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of California, San Diego, 9500 Gilman Drive, La Jolla, California 92093-0356, USA.