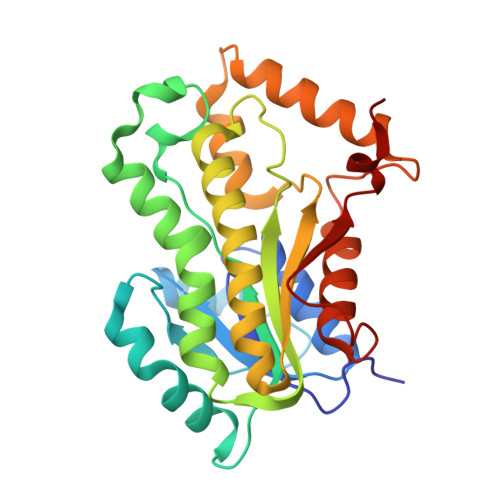
Triclosan Derivatives: Towards Potent Inhibitors of Drug-Sensitive and Drug-Resistant Mycobacterium tuberculosis.
Freundlich, J.S., Wang, F., Vilcheze, C., Gulten, G., Langley, R., Schiehser, G.A., Jacobus, D.P., Jacobs, W.R., Sacchettini, J.C.(2009) ChemMedChem 4: 241-248
- PubMed: 19130456
- DOI: https://doi.org/10.1002/cmdc.200800261
- Primary Citation of Related Structures:
3FNE, 3FNF, 3FNG, 3FNH - PubMed Abstract:
Triclosan has been previously shown to inhibit InhA, an essential enoyl acyl carrier protein reductase involved in mycolic acid biosynthesis, the inhibition of which leads to the lysis of Mycobacterium tuberculosis. Using a structure-based drug design approach, a series of 5-substituted triclosan derivatives was developed. Two groups of derivatives with alkyl and aryl substituents, respectively, were identified with dramatically enhanced potency against purified InhA. The most efficacious inhibitor displayed an IC(50) value of 21 nM, which was 50-fold more potent than triclosan. X-ray crystal structures of InhA in complex with four triclosan derivatives revealed the structural basis for the inhibitory activity. Six selected triclosan derivatives were tested against isoniazid-sensitive and resistant strains of M. tuberculosis. Among those, the best inhibitor had an MIC value of 4.7 microg mL(-1) (13 microM), which represents a tenfold improvement over the bacteriocidal activity of triclosan. A subset of these triclosan analogues was more potent than isoniazid against two isoniazid-resistant M. tuberculosis strains, demonstrating the significant potential for structure-based design in the development of next generation antitubercular drugs.
Organizational Affiliation:
Texas A&M University, Department of Biochemistry and Biophysics, College Station, 77843-2128, USA.