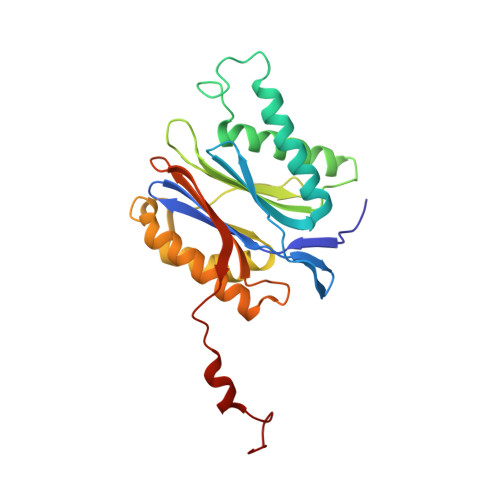
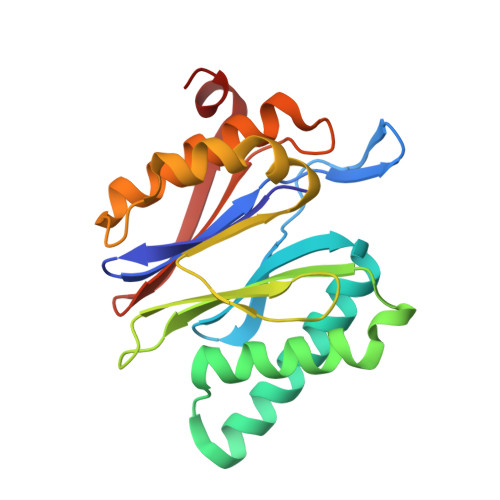
Inhibitor-binding mode of homobelactosin C to proteasomes: new insights into class I MHC ligand generation
Groll, M., Larionov, O.V., Huber, R., de Meijere, A.(2006) Proc Natl Acad Sci U S A 103: 4576-4579
- PubMed: 16537370
- DOI: https://doi.org/10.1073/pnas.0600647103
- Primary Citation of Related Structures:
3E47 - PubMed Abstract:
Most class I MHC ligands are generated from the vast majority of cellular proteins by proteolysis within the ubiquitin-proteasome pathway and are presented on the cell surface by MHC class I molecules. Here, we present the crystallographic analysis of yeast 20S proteasome in complex with the inhibitor homobelactosin C. The structure reveals a unique inhibitor-binding mode and provides information about the composition of proteasomal primed substrate-binding sites. IFN-gamma inducible substitution of proteasomal constitutive subunits by immunosubunits modulates characteristics of generated peptides, thus producing fragments with higher preference for binding to MHC class I molecules. The structural data for the proteasome:homobelactosin C complex provide an explanation for involvement of immunosubunits in antigen generation and open perspectives for rational design of ligands, inhibiting exclusively constitutive proteasomes or immunoproteasomes.
Organizational Affiliation:
Ludwig Maximilians Universität, Adolf Butenandt Institut, Butenandtstrasse 5, Gebäude B, D-81377 Munich, Germany. mgroll@med.uni-muenchen.de