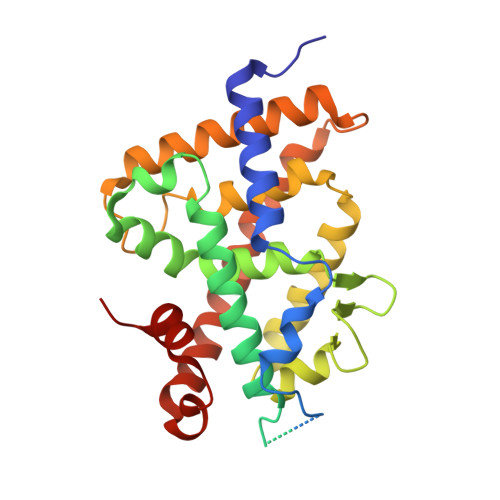
Superagonistic fluorinated vitamin D3 analogs stabilize helix 12 of the vitamin D receptor.
Eelen, G., Valle, N., Sato, Y., Rochel, N., Verlinden, L., De Clercq, P., Moras, D., Bouillon, R., Munoz, A., Verstuyf, A.(2008) Chem Biol 15: 1029-1034
- PubMed: 18940664
- DOI: https://doi.org/10.1016/j.chembiol.2008.08.008
- Primary Citation of Related Structures:
3DR1 - PubMed Abstract:
Side chain fluorination is often used to make analogs of 1,25-dihydroxyvitamin D3 [1,25(OH)2D3] resistant to degradation by 24-hydroxylase. The fluorinated nonsteroidal analogs CD578, WU515, and WY1113 have an increased prodifferentiating action on SW480-ADH colon cancer cells, which correlated with stronger induction of vitamin D receptor (VDR)-coactivator interactions and stronger repression of beta-catenin/TCF activity. Cocrystallization of analog CD578 with the zebrafish (z)VDR and an SRC-1 coactivator peptide showed that the fluorine atoms of CD578 make additional contacts with Val444 and Phe448 of activation helix 12 (H12) of the zVDR and with Leu440 of the H11-H12 loop. Consequently, the SRC-1 peptide makes more contacts with the VDR-CD578 complex than with the VDR-1,25(OH)2D3 complex. These data show that fluorination not only affects degradation of an analog but can also have direct effects on H12 stabilization.
Organizational Affiliation:
LEGENDO, K.U. Leuven, B-3000 Leuven, Belgium.