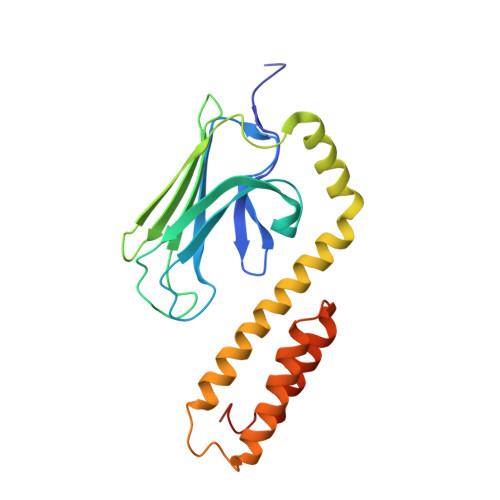
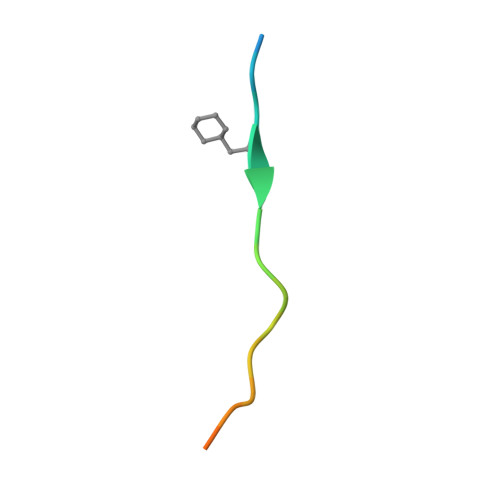
Allosteric coupling between the lid and interdomain linker in DnaK revealed by inhibitor binding studies.
Liebscher, M., Roujeinikova, A.(2009) J Bacteriol 191: 1456-1462
- PubMed: 19103929
- DOI: https://doi.org/10.1128/JB.01131-08
- Primary Citation of Related Structures:
3DPO, 3DPP, 3DPQ - PubMed Abstract:
The molecular chaperone DnaK assists protein folding and refolding, translocation across membranes, and regulation of the heat shock response. In Escherichia coli, the protein is a target for insect-derived antimicrobial peptides, pyrrhocoricins. We present here the X-ray crystallographic analysis of the E. coli DnaK substrate-binding domain in complex with pyrrhocoricin-derived peptide inhibitors. The structures show that pyrrhocoricins act as site-specific, dual-mode (competitive and allosteric) inhibitors, occupying the substrate-binding tunnel and disrupting the latch between the lid and the beta-sandwich. Our structural analysis revealed an allosteric coupling between the movements of the lid and the interdomain linker, identifying a previously unknown mechanism of the lid-mediated regulation of the chaperone cycle.
Organizational Affiliation:
Martin Luther University Halle-Wittenberg, Saale, Germany.