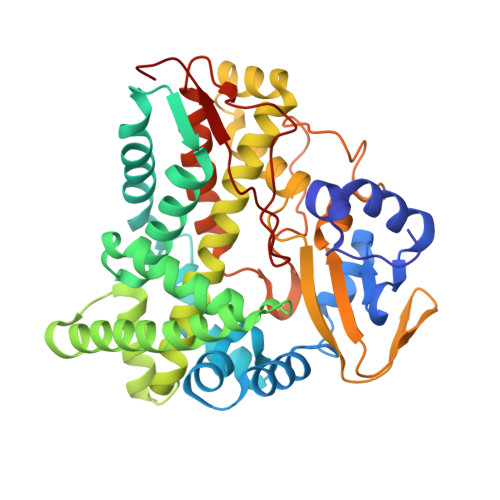
Structural characterization of CalO2: a putative orsellinic acid P450 oxidase in the calicheamicin biosynthetic pathway.
McCoy, J.G., Johnson, H.D., Singh, S., Bingman, C.A., Lei, I.K., Thorson, J.S., Phillips Jr., G.N.(2009) Proteins 74: 50-60
- PubMed: 18561189
- DOI: https://doi.org/10.1002/prot.22131
- Primary Citation of Related Structures:
3BUJ - PubMed Abstract:
Although bacterial iterative Type I polyketide synthases are now known to participate in the biosynthesis of a small set of diverse natural products, the subsequent downstream modification of the resulting polyketide products remains poorly understood. Toward this goal, we report the X-ray structure determination at 2.5 A resolution and preliminary characterization of the putative orsellenic acid P450 oxidase (CalO2) involved in calicheamicin biosynthesis. These studies represent the first crystal structure for a P450 involved in modifying a bacterial iterative Type I polyketide product and suggest the CalO2-catalyzed step may occur after CalO3-catalyzed iodination and may also require a coenzyme A- (CoA) or acyl carrier protein- (ACP) bound substrate. Docking studies also reveal a putative docking site within CalO2 for the CLM orsellinic acid synthase (CalO5) ACP domain which involves a well-ordered helix along the CalO2 active site cavity that is unique compared with other P450 structures.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin-Madison, Madison, Wisconsin 53706-1544, USA.