Structural, Biochemical, and Phylogenetic Analyses Suggest That Indole-3-Acetic Acid Methyltransferase Is an Evolutionarily Ancient Member of the SABATH Family.
Zhao, N., Ferrer, J.L., Ross, J., Guan, J., Yang, Y., Pichersky, E., Noel, J.P., Chen, F.(2008) Plant Physiol 146: 455-467
- PubMed: 18162595
- DOI: https://doi.org/10.1104/pp.107.110049
- Primary Citation of Related Structures:
3B5I - PubMed Abstract:
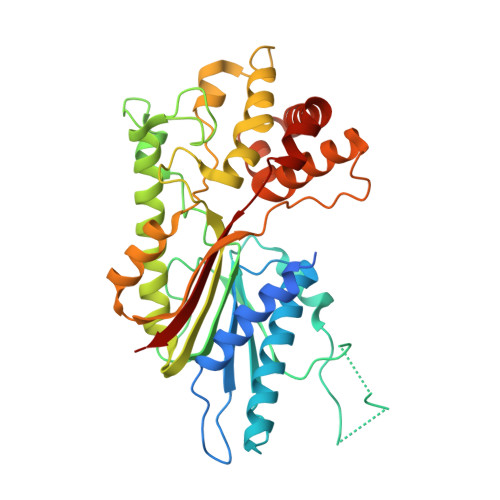
The plant SABATH protein family encompasses a group of related small-molecule methyltransferases (MTs) that catalyze the S-adenosyl-L-methionine-dependent methylation of natural chemicals encompassing widely divergent structures. Indole-3-acetic acid (IAA) methyltransferase (IAMT) is a member of the SABATH family that modulates IAA homeostasis in plant tissues through methylation of IAA's free carboxyl group. The crystal structure of Arabidopsis (Arabidopsis thaliana) IAMT (AtIAMT1) was determined and refined to 2.75 A resolution. The overall tertiary and quaternary structures closely resemble the two-domain bilobed monomer and the dimeric arrangement, respectively, previously observed for the related salicylic acid carboxyl methyltransferase from Clarkia breweri (CbSAMT). To further our understanding of the biological function and evolution of SABATHs, especially of IAMT, we analyzed the SABATH gene family in the rice (Oryza sativa) genome. Forty-one OsSABATH genes were identified. Expression analysis showed that more than one-half of the OsSABATH genes were transcribed in one or multiple organs. The OsSABATH gene most similar to AtIAMT1 is OsSABATH4. Escherichia coli-expressed OsSABATH4 protein displayed the highest level of catalytic activity toward IAA and was therefore named OsIAMT1. OsIAMT1 exhibited kinetic properties similar to AtIAMT1 and poplar IAMT (PtIAMT1). Structural modeling of OsIAMT1 and PtIAMT1 using the experimentally determined structure of AtIAMT1 reported here as a template revealed conserved structural features of IAMTs within the active-site cavity that are divergent from functionally distinct members of the SABATH family, such as CbSAMT. Phylogenetic analysis revealed that IAMTs from Arabidopsis, rice, and poplar (Populus spp.) form a monophyletic group. Thus, structural, biochemical, and phylogenetic evidence supports the hypothesis that IAMT is an evolutionarily ancient member of the SABATH family likely to play a critical role in IAA homeostasis across a wide range of plants.
Organizational Affiliation:
Department of Plant Sciences, University of Tennessee, Knoxville, Tennessee 37996, USA.