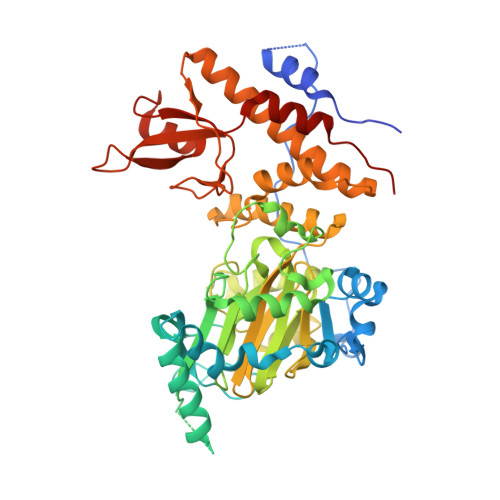
Structural basis for histone H3 Lys 27 demethylation by UTX/KDM6A
Sengoku, T., Yokoyama, S.(2011) Genes Dev 25: 2266-2277
- PubMed: 22002947
- DOI: https://doi.org/10.1101/gad.172296.111
- Primary Citation of Related Structures:
3AVR, 3AVS - PubMed Abstract:
Tri- and dimethylations of histone H3K9 (H3K9me3/2) and H3K27 (H3K27me3/2), both situated in the "A-R-Kme-S" sequence motif, mediate transcriptional repression of distinct genomic regions. H3K9me3/2 mainly governs constitutive heterochromatin formation, while H3K27me3/2 represses key developmental genes. The mechanisms by which histone-modifying enzymes selectively regulate the methylation states of H3K9 and H3K27 are poorly understood. Here we report the crystal structures of the catalytic fragment of UTX/KDM6A, an H3K27me3/2-specific demethylase, in the free and H3 peptide-bound forms. The catalytic jumonji domain binds H3 residues 25-33, recognizing H3R26, H3A29, and H3P30 in a sequence-specific manner, in addition to H3K27me3 in the catalytic pocket. A novel zinc-binding domain, conserved within the KDM6 family, binds residues 17-21 of H3. The zinc-binding domain changes its conformation upon H3 binding, and thereby recognizes the H3L20 side chain via a hydrophobic patch on its surface, which is inaccessible in the H3-free form. Mutational analyses showed that H3R17, H3L20, H3R26, H3A29, H3P30, and H3T32 are each important for demethylation. No other methyllysines in the histone tails have the same set of residues at the corresponding positions. Thus, we clarified how UTX discriminates H3K27me3/2 from the other methyllysines with distinct roles, including the near-cognate H3K9me3/2, in histones.
Organizational Affiliation:
RIKEN Systems and Structural Biology Center, Tsurumi, Yokohama, Japan.